217
218
Summary
The data presented below will show several major conclusions:
(a) The male Basques living today have rather recent
roots of less than four thousand years ago, contrary to
the legend that proposes they lived some 30 thousand
years ago.
(b) There is no justification in the results of a
“Ukrainian refuge” for the R1a1 ancient population
allegedly 15,000 years ago; instead, evidence has been
obtained that the oldest R1a1 lived circa 20,000 years
before the present (ybp) in South Siberia. There are two
sets of data and these provide ages of 21,000±3,000 ybp
and 19,625±2,800 ybp, calculated by two different
methods, and 11,650±1,550 years ago they appeared in the
Balkans (Serbia, Kosovo, Bosnia, Macedonia).
(c) Except the South Siberian and Balkans populations,
present-day bearers of R1a1 across Western and Eastern
Europe have common ancestors who lived between
3550 and 4750 years ago (the “youngest” in Scotland,
Ireland and Sweden, the “oldest” in Russia (4750±500
ybp) and Germany (4,700±520 ybp),
(d) There are two different groups of Indian R1a1
haplotypes; one shows a good match with the Russian
Slavic R1a1 group, having a common ancestor several
hundred years “younger” than the Russian R1a common
ancestor (4,050±500 vs. 4,750±500 ybp). This
supports the idea that a proto-Slavic migration to India
as Aryans occurred (mentioned in classic ancient Indian
literature) around 3600 ybp. The other Indian R1a
population is significantly older, with a common ancestor
living 7,125±950 ybp; they could have migrated
from South Siberia to South India.
(e) South India Chenchu R1a1 match the current Russian
Slavic R1a1 haplotypes, and the Chenchu R1a
common ancestor appeared some 3200±1900 ybp, apparently
after the R1a1 migration from the North to
India. Another Chenchu R1a1 lineage originated about
350±350 ybp, around the 17th century CE.
(f) The so-called Cohen Modal Haplotype in Haplogroup
J1 originated 9,000±1,400 years ago, if all related
J1 haplotypes are considered. About 4,000±520 ybp it
appeared in the proto-Jewish population, and
1,050±190 years ago (if to consider only CMH) or
1,400±260 years ago (if to consider only Jewish J1
population) split a “recent CMH” lineage.
(g) another so-called CMH, of Haplogroup J2, appeared
in the Jewish population 1,375±300 years ago,
(h)The South African Lemba population of Haplogroup
J has nothing to do with ancient Jewish patriarchs, since
the haplogroup appears to have penetrated the Lemba
population some 625±200 ybp, around the 14th century
CE.
(i) Native American Haplogroup Q1a3a contains at
least six lineages, the oldest of which originated
16,000±3,300 years ago, in accord with archaeological
data.
The methods used in the present study are described in
the companion article.
Results and Discussion: Eurasian R1a1 Populations
The “mapping” of the enormous territory from the Atlantic through Russia and India to the
Pacific, and from Scandinavia to the Arabian Peninsula, reveals that Haplogroup R1a men
are marked with practically the same ancestral haplotype, which is about 4,500 - 4,700
years “old.” through much of its geographic range.
Exceptions in Europe are found only in the Balkans (Serbia, Kosovo, Macedonia, Bosnia),
where the common ancestor is significantly more ancient, about 11,650±1,550 ybp, and in
the Irish, Scottish and Swedish R1a1 populations, which have a significantly “younger”
common ancestor, some thousands years “younger” compared with the Russian, the German,
and the Poland R1a1 populations. These geographic patterns will be explored below in this
section. The haplogroup name R1a1 is used here to mean Haplogroup R-M17, because that is
the meaning in nearly all of the referenced articles.
The entire map of base (ancestral) haplotypes and their mutations, as well as “ages” of
common ancestors of R1a1 haplotypes in Europe, Asia, and the Middle East show that
approximately six thousand years ago bearers of R1a1 haplogroup started to migrate from
the Balkans in all directions, spreading their haplotypes. A recent excavation of 4,600
year-old R1a1 haplotypes (Haak et al., 2008) revealed their almost exact match to
present day R1a1 haplotypes, as it is shown below.
Significantly, the oldest R1a population appears to be in Southern Siberia.
England and Ireland R1a1 haplotypes
The 57 R1a 25-marker haplotype series of English origin (YSearch database) contains ten
haplotypes that belong to a DYS388=10 series and was analyzed separately.
The remaining 47 haplotypes contain 304 mutations compared to the base haplotype shown
below, which corresponds to 4,125±475 years to a common ancestor in the 95% confidence
interval. The respective haplotype tree is shown in Fig. 3.
225
| Fig. 3 English origin |
Fig.4 Ireland origin |
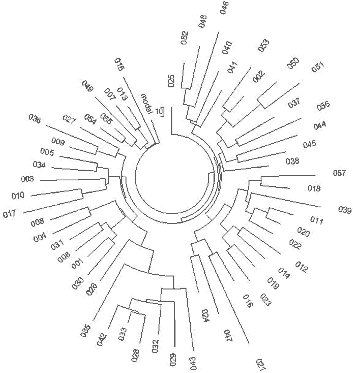 |
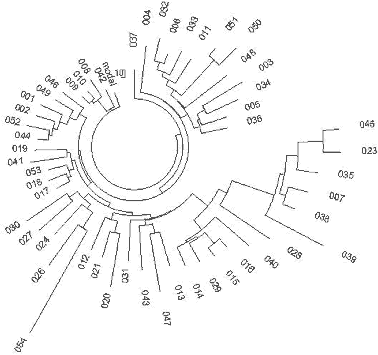 |
The 52
haplotype series of Ireland origin, all with 25 markers (YSearch database) contains 12
haplotypes which belong to a DYS388=10 distinct series as shown in Fig. 4, and was analyzed
separately. The remaining 40 haplotypes contain 244 mutations compared to the base
haplotype, shown below, which corresponds to 3,850±460 years to a common ancestor.
Thus, R1a1 haplotypes sampled on the British Isles point to English and Irish common
ancestors who lived 4,125±475 and 3,850±460 years ago. The English base (ancestral)
haplotype is as follows
13-25-15-10-11-14-12-12-10-13-11-30-15-9-10-11-11- 24-14
20-32-12-15-15-16
and the Irish one:
13-25-15- -11-14-12-12-10-13-11-30-15-9-10-11-11-
-14 20-32-12-15-15-16
An apparent difference in two alleles between the British and Irish
ancestral haplotypes is in fact fairly insignificant, since the respective average
alleles are equal to 10.51 and 10.73, and 23.98 and 23.55, respectively.
Hence, their ancestral haplotypes are practically the same, within approximately one
mutational difference.
A DYS388=10 Subfamily of North-Western European R1a1 Haplotypes
About 20% of both English and Irish R1a haplotypes have a mutated allele in eighth
position in the FTDNA format (DYS388=12à10), with a common ancestor of that population
who lived 3,575±450 years ago (172 mutations in the 30 25-marker haplotypes with
DYS388=10). 61 of these haplotypes were pooled from a number of European populations (
Fig. 5),
and the tree splits into a relatively younger branch on the left, and the “older” branch
on the lower right-hand side.
226
| Fig. 5 Subfamily of North-Western European R1a1 Haplotype with DYS388=10 |
Fig. 6 European R1a1 Haplotypes |
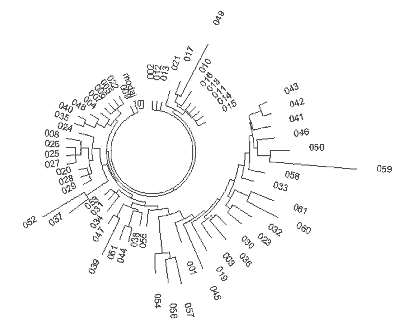 |
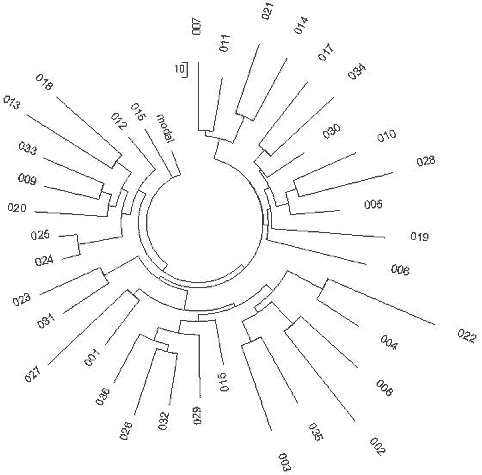 |
This DYS388=10 mutation is observed in northern and western Europe, mainly in
England, Ireland, Norway, and to a much lesser degree in Sweden, Denmark, Netherlands and
Germany. In areas further east and south that mutation is practically absent.
31 haplotypes on the left-hand side and on the top of the tree (Fig. 5) contain collectively
86 mutations from the base haplotype:
13-25-15-10-11-14-12-10-10-13-11-30-15-9-10-11-11-
25-14-19-32-12-14-14-17
which corresponds to 1625±240 years to the common ancestor. It is
a rather recent common ancestor, who lived around the 4th century CE. The closeness of
the branch to the trunk of the tree in also points to the rather recent origin of the
lineage.
The older 30-haplotype branch in the tree provides with the following base DYS388=10
haplotype:
13-25-16-10-11-14-12-10-10-13-11-30-15-9-10-11-11- 24-14-19-32-12-14-15-16
These 30 haplotypes contain 172 mutations, for which the linear method gives 3575±450
years to the common ancestor.
These two DYS388=10 base haplotypes differ from each other by less than four mutations,
which brings their common ancestor to about 3500 ybp. It is very likely that it is the
same common ancestor as that of the right-hand branch in Fig. 5.
The upper, “older” base haplotype differs by six mutations on average from the DYS388=12
base haplotypes from the same area (see the English and Irish R1a1 base haplotypes,
above). This brings common ancestor to about 5,700±600 ybp.
This common ancestor of both the DYS388=12 and DYS388=10 populations lived presumably in
the Balkans (see below) since the Balkan population is the only European R1a1 population
that is that old, for almost two thousand years before bearers of that mutation arrived
to northern and western Europe some 4000 ybp (DYS388=12) and about 3600 ybp (DYS388=10).
This mutation has continued to be passed down through the generations to the present
time.
227
Scotland R1a1 Haplotypes
A set of 29 R1a1 25-marker haplotypes from Scotland have the same ancestral haplotype
as those in England and Ireland:
13-25-15-11-11-14-12-12-10-13-11-30-15-9-10-11-11- 24-14
20-32-12-15-15-16
This set of haplotypes contained 164 mutations, which gives 3550±450
years to the common ancestor.
Germany R1a1 Haplotypes
A 67-haplotype series with 25 markers from Germany revealed the following ancestral
haplotype:
13-25-16-10-11-14-12-12-10-13-11-30-15-9-10-11-11- 24-14 20-32-12-15-15-16
There is an apparent mutation in the third allele from the left (in bold) compared with
the Isles ancestral R1a1 haplotypes, however, the average value equals to 15.48 for
England, 15.33 for Ireland, 15.26 for Scotland, and 15.84 for Germany, so there is only a
small difference between these populations. The 67 haplotypes contain 488 mutations, or
0.291±0.013 mutations per marker on average. This value corresponds to 4,700±520 years
from a common ancestor in the German territory.
These results are supported by the very recent data found during excavation of remains
from several males near Eulau, Germany. Tissue samples from one of them was derived for
the SNP, SRY10831.2 (Haak et al., 2008), so it was from Haplogroup R1a1 and probably
R1a1a as well (the SNPs defining R1a1a were not tested). The skeletons were dated to
4,600 ybp using strontium isotope analysis. The 4600 year old remains yielded some DNA
and a few STRs were detected, yielding the following partial haplotype:
13(14)-25-16-11-11-14-X-Y-10-13-Z-30-15
These haplotypes very closely resemble the above
R1a1 ancestral haplotype in Germany both in the structure and in the dating (4,700±520
and 4,600 ybp).
Norway and Sweden R1a1 Haplotypes
The ancestral R1a1 haplotypes for the Norwegian and Swedish groups are almost the same,
and both are similar to the German 25-marker ancestral haplotype. Their 16-and
19-haplotype sets (after DYS388=10 haplotypes were removed, five and one, respectively)
contain on average 0.218±0.023 and 0.242±0.023 mutations per marker, respectively, which
gives a result of 3,375±490 and 3,825±520 years back to their common ancestors. This is likely the same time span within the error margin.
Polish, Czech, and Slovak R1a1 Haplotypes
The ancestral haplotypes for the Polish, Czech, and Slovak groups are very similar to
each other, having only one insignificant difference in DYS439 (shown in bold). The average value is 10.43 on DYS439 in the Polish group, 10.63 in the Czech and Slovak
combined groups. The base haplotype for the 44 Polish haplotypes is:.
13-25-16-10-11-14-12-12- -13-11-30-16-9-10-11-11- 23-14-20-32-12-15-15-16
and for the 27 Czech and Slovak haplotypes it is:
13-25-16-10-11-14-12-12- 13-11-30-16-9-10-11-11- 23-14-20-32-12-15-15-16
The difference
in these haplotypes is really only 0.2 mutations - the alleles are just rounded in
opposite directions.
The 44 Polish haplotypes have 310 mutations among them, and there are 175 mutations in
the 27 Czech-Slovak haplotypes, resulting in 0.282±0.016 and 0.259±0.020 mutations per
marker on average, respectively. These values correspond to 4,550±520 and 4,125±430 years back to their common ancestors,
respectively, which is very similar to the German TSCA.
The Remaining European R1a1 Haplotypes
Many European countries are represented by just a few 25-marker haplotypes in the
databases. 36 R1a1 haplotypes were collected from the YSearch database where the
ancestral origin of the paternal line was from Denmark, Netherlands, Switzerland,
Iceland, Belgium, France, Italy, Lithuania, Romania, Albania, Montenegro, Slovenia,
Croatia, Spain, Greece, Bulgaria and Moldavia. The respective haplotype tree is shown in
Fig. 6.
The tree does not show any noticeable anomalies and points at just one common ancestor
for all 36 individuals, who had the following base haplotype:
13-25-16-10-11-14-12-12-10-13-11-30-15-9-10-11-11- 24-14-20-32-12-15-15-16
The ancestral
haplotype match is exactly the same as those in Germany, Russia (see below), and has
quite insignificant deviations from all other ancestral haplotypes considered above,
within fractions of mutational differences. All the 36 individuals have 248 mutations in
their 25-marker haplotypes, which corresponds to 4,425±520 years to the common ancestor.
This is quite a common value for European R1a1 population.
Russia and Ukraine R1a1 Haplotypes
The haplotype tree containing 110 of 25-marker haplotypes collected over 10 time zones
from the Western Ukraine to the Pacific Ocean and from the northern tundra to Central
Asia (Tadzhikistan and Kyrgyzstan) is shown in Fig. 7.
228
| Fig. 7 Russia and Ukraine R1a1 Haplotype |
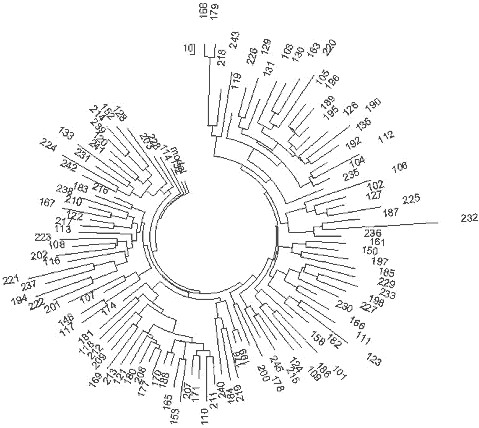 |
The ancestral (base) haplotype for the haplotype tree is
13-25-16-11-11-14-12-12-10-13-11-30-15-9-10-11-11- 24-14-20-32-12-15-15-16
It is almost
exactly the same ancestral haplotype as that in Germany. The average value on DYS391 is
10.53 in the Russian haplotypes, and was rounded to 11, and the base haplotype has only
one insignificant deviation from the ancestral haplotype in England, which has the third
allele (DYS19) 15.48, which in Russia/Ukraine it is 15.83 (calculated from one DYS19=14,
32 DYS19=15, 62 DYS19=16, and 15 DYS19=17). There are similar insignificant deviations
with the Polish and Czechoslovakian base haplotypes, at DYS458 and DYS447, respectively,
within 0.2-0.5 mutations. This indicates a small mutational difference between their
common ancestors.
The 110 Russian/Ukranian haplotypes contain 804 mutations from the base haplotype, or
0.292±0.010 mutations per marker, resulting in 4,750±500 years from a common ancestor.
The degree of asymmetry of this series of haplotypes is exactly 0.50, and does not affect
the calculations.
229
R1a1 haplotypes of individuals who considered themselves of Ukrainian and Russian origin,
present a practically random geographic sample. For example, the haplotype tree contains two local
Central Asian haplotypes (a Tadzhik and a Kyrgyz, haplotypes 133 and 127, respectively),
as well as a local of a Caucasian Mountains Karachaev tribe (haplotype 166), though the
male ancestry of the last one is unknown beyond his present tribal affiliation. They did
not show any unusual deviations from other R1a1 haplotypes. Apparently, they are derived
from the same common ancestor as are all other individuals of the R1a1 set.
The literature frequently refers to a statement that R1a1- M17 originated from a “refuge”
in the present Ukraine about 15,000 years ago, following the Last Glacial Maximum. This
statement was never substantiated by any actual data related to haplotypes and
haplogroups.
It is just repeated over and over through a relay of references to references. The oldest
reference is apparently that of Semino et al. (2000) which states that “this scenario is
supported by the finding that the maximum variation for microsatellites linked to Eu19
[R1a1] is found in Ukraine” (Santachiara-Benerecetti, unpublished data). Now we know that
this statement is incorrect. No calculations were provided in (Semino et al, 2000) or
elsewhere which would explain the dating of 15,000 years.
Then, a paper by Wells et al. (2001) states “M17, a descendant of M173, is apparently
much younger, with an inferred age of ~ 15,000 years.” No actual data or calculations are
provided. The subsequent sentence in the paper says, “It must be noted that these age
estimates are dependent on many, possibly invalid, assumptions about mutational
processes and population structure.” This sentence has turned out to be valid in the
sense that the estimate was inaccurate and overestimated by about 300% from the results
obtained here. However, see the results below for the Chinese and southern Siberian
haplotypes, below.
230
A more detailed consideration of R1a1 haplotypes in the Eastern
European Plain and across Europe
and Eurasia in general has shown that R1a1 haplogroup appeared in Europe between 12 and
10 thousand years before present, right after the Last Glacial Maximum, and after about
6,000 ybp had populated Europe, though, probably, with low density. After 4,500 ybp R1a1
practically disappeared from Europe, incidentally, along with I1. Maybe more
incidentally, it corresponded with the time period of populating of Europe with R1b1b2.
Only those R1a1 who migrated to the Eastern European Plain from Europe around 6,000-5,000
ybp, stayed. They had expanded to the East, established on their way a number of
archaeological cultures, including the Andronovo culture, which has embraced Northern
Kazakhstan, Central Asia and South Ural and Western Siberia, and about 3600 ybp they
migrated to India and Iran as the Aryans. Those who left behind, on the Eastern European
Plain,
re-populated Europe between 3200 and 2500 years bp, and stayed mainly in the Eastern
Europe (present-day Poland, Germany, Czech, Slovak, etc. regions). Among them were carriers of the
newly discovered R1a1-M458 subclade (Underhill et al, 2009).
|
This conclusion is consistent with the conclusion of Mario Alinei
Paleolithic Continuity Theory on Slavic presence in post-glacial Balkans, and with A. Dybo comparative analysis of Pra-IE and Pra-Altaic linguistic groups
Pra-Altaian World |
231
India R1a1 Haplotypes
The YSearch database contains 22 of the 25-marker R1a1 haplotypes from India, including a
few haplotypes from Pakistan and Sri Lanka. Their ancestral haplotype follows:
13-25-16-10-11-14-12-12-10-13-11-30-16-9-10-11-11- 24-14-20-32-12-15-15-16
The only one
apparent deviation in DYS458 (shown in bold) is related to an average alleles equal to
16.05 in Indian haplotypes, and 15.28 in Russian ones.
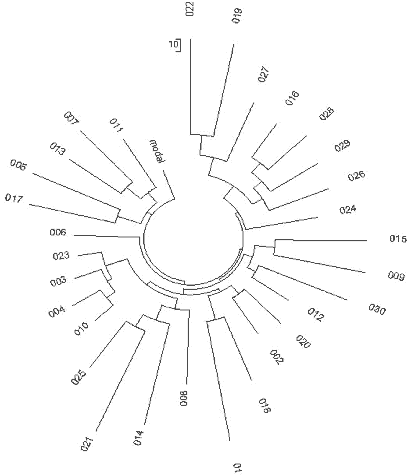
The India (Regional) Y-project at FTDNA (Rutledge, 2009) contains 15 haplotypes with 25
markers, eight of which are not listed in the YSearch database. Combined with others from
Ysearch, a set of 30 haplotypes is available. Their ancestral haplotype is exactly the
same as shown above. Fig. 8 shows the respective haplotypes tree.
All 30 Indian R1a1 haplotypes
contain 191 mutations, that is 0.255±0.018 mutations per marker. It is statistically
lower (with 95% confidence interval) than 0.292±0.014 mutations per marker in the Russian
haplotypes and corresponds to 4,050±500 years from a common ancestor of the Indian
haplotypes, compared to 4,750±500 years for the ancient “Russian” TSCA.
Archaeological studies have been conducted since the 1990’s in the South Ural’s Arkaim
settlement and have revealed that the settlement was abandoned 3,600 years ago. The
population apparently moved to northern India. That population belonged to Andronovo
archaeological culture. Excavations of some sites of Andronovo culture, the oldest dating
between 3,800 and 3,400 ybp, showed that nine inhabitants out of ten shared the R1a1
haplogroup and haplotypes (Bouakaze et al., 2007; Keyser et al, 2009). The base haplotype
is as follows:
13-25(24)-16(17)-11-11-14-X-Y-10-13(14)-11-31(32)
In this example,
alleles that have not been assessed are replaced with letters. One can see that the
ancient R1a1 haplotype closely resembles the Russian (as well as the other R1a1)
ancestral haplotypes.
232
In a recent paper (Keyser et al, 2009) the authors have extended the earlier analysis
and determined 17-marker haplotypes for 10 Siberian individuals assigned to Andronovo, Tagar, and Tashtyk archaeological cultures
(See Keyser et al, 2009
Sibirian Kurgan Y- and mt-DNA
 .
.
| Fig. 8 India R1a1 Haplotype |
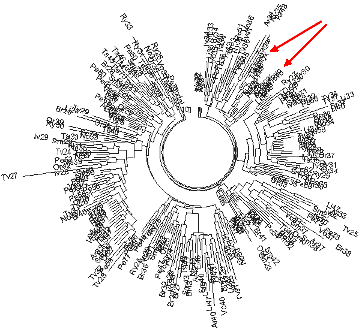
Figure 9. The 17-marker haplotype tree for R1a1 haplotypes of 262 ethnic Russians, most from Roewer et al (2008),
to which seven ancient (excavated) Andronovo, Tagar, and Tashtyk haplotypes (Keyser et al, 2009) were added (their positions are indicated with the arrows; two more haplotypes
were identical with two from the selection. See also Figure 10. |
 |
 |
| |
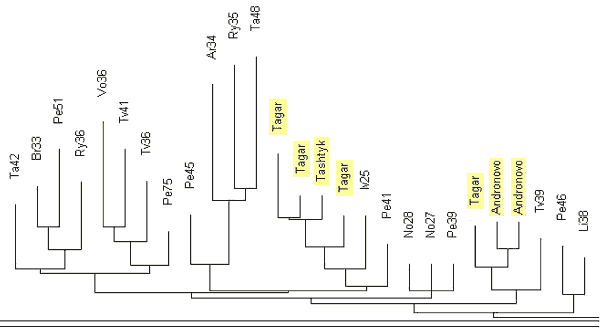
Figure 10. A fragment of the 17-marker haplotype tree for R1a1 haplotypes of ethnic Russians,
to which seven ancient (excavated) Andronovo, Tagar, and Tashtyk haplotypes were added
(Figure 9). Two-letter designations for the Russian regions are as follows: |
|
| Ar-Archangelsk |
Iv-Ivanovo |
No-Novgorod |
Ry-Ryazan |
Tv-Tver |
| Br-Bryansk |
Li-Lipezk |
Pe-Penza |
Ta-Tambov |
Vo-Vologda |
 |
Nine of them shared the R1a1 haplogroup (one was of Haplogroup CxC3). The authors have
reported that “none of the Y-STR haplotypes perfectly matched those included in the
databases,” and for some of those haplotypes “even the search based on the 9-loci minimal
haplotype was fruitless.” However, as shows, all of the haplotypes nicely fit to the
17-marker haplotype tree of 252 Russian R1a1 haplotypes (with a common ancestor of
4,750±490 ybp, calculated using these 17-marker haplotypes [Klyosov, 2009b]), a list of
which was recently published (Roewer et al, 2008). All seven Andronovo, Tagar and Tashtyk
haplotypes (the other two were incomplete) are located in the upper right hand side corner
in Fig. 9, and the respective branch on the tree is shown in Fig. 10.
As one can see, the ancient R1a1 haplotypes excavated in Siberia, are comfortably
located on the tree next to the haplotypes from the Russian cities and regions named in
the legend to Fig. 10.
The above data provide rather strong evidence that the R1a1 tribe migrated from Europe to
the East between 5,000 and 3,600 ybp. The pattern of this migration is exhibited as
follows:
1) the descendants who live today share a common ancestor of 4,725±520 ybp,
2)
the Andronovo (and the others) archaeological complex of cultures in North Kazakhstan and
South and Western Siberia dates 4,300 to 3,500 ybp, and it revealed several R1a1
excavated haplogroups,
3) they reached the South Ural region some 4,000 ybp, which is
where they built Arkaim, Sintashta (contemporary names), and the so called “a country of
towns” in the South Ural region around 3,800 ybp,
4) by 3,600 ybp they abandoned the area
and moved to India under the name of Aryans. The Indian R1a1 common ancestor of 4,050±500
ybp chronologically corresponds to these events. Currently, some 16% of the Indian
population, that is about 100 millions males, and the majority of the upper castes, are
members of Haplogroup R1a1 (Sengupta et al, 2006; Sharma et al, 2009).
Since we have mentioned Haplogroup R1a1 in India and in the archaeological cultures north
of it, it is worthwhile to consider the question of where and when R1a1 haplotypes
appeared in India. On the one hand, it is rather obvious from the above, that
R1a1
haplotypes were brought to India around 3500 ybp from what is now Russia. The R1a1
bearers, known later as the Aryans, brought to India not only their haplotypes and the
haplogroup, but also their language, thereby closing the loop, or building the linguistic
and cultural bridge between India (and Iran) and Europe, and possibly creating the
Indo-European family of languages. On the other hand, there is evidence that some Indian
R1a1 haplotypes show a high variance, exceeding that in Europe (Kivisild et al, 2003;
Sengupta et al, 2006; Sharma et al, 2009; Thanseem et al, 2009; Fornarino et al, 2009),
thereby antedating the 4000-year-old migration from the north. However, there is a way to resolve the
conundrum.
233
It seems that there are two quite distinct sources of Haplogroup R1a1 in India. One,
indeed, was probably brought from the north by the Aryans. However, the most ancient
source of R1a1 haplotypes appears to be provided by people who now live in China.
In an article by Bittles et al (2007) entitled “Physical anthropology and ethnicity in
Asia: the transition from anthropometry to genome-based studies” a list of frequencies of
Haplogroup R1a1 is given for a number of Chinese populations, however, haplotypes were
not provided. The corresponding author, Professor Alan H. Bittles, kindly sent me the following list of 31 five-marker haplotypes (presented here
in the format DYS19, 388, 389-1, 389-2, 393), in Table 3. The haplotype tree is shown in
Fig. 11.
Table 3
31 Five-Marker R1a1 Haplotypes from China
(DYS 19, 388, 389-1, 389-2, 393) |
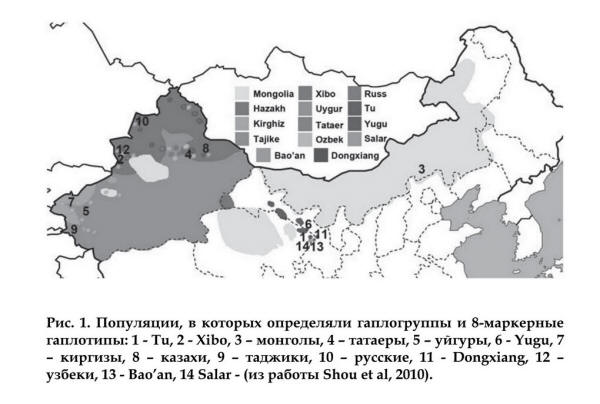
Populations in Shou et al, 2010: 1 Tu (Monguor/Tu=Tibetans/Uigurs?)), 2 Xibo
(Manchu/Türkic?), 3 Mongols, 4 Tatars, 5 Uigurs, 6 Yugu (Yugur=Uigur), 7 Kirgiz, 8 Kazakh, 9 Tajik, 10 Russian, 11
Dongxiang, 12 Uzbeks, 13 Bao'an (Uigur), 14 Salar |
| 17 12 13 30 13 |
14 12 13 30 12 |
14 12 12 31 13 |
 |
| 14 13 13 32 13 |
14 12 13 30 13 |
17 12 13 32 13 |
| 14 12 13 32 13 |
17 12 13 30 13 |
17 12 13 31 13 |
| 17 12 12 28 13 |
14 12 13 30 13 |
14 12 12 29 13 |
| 17 12 14 31 13 |
14 12 12 28 13 |
14 13 14 30 13 |
| 14 12 12 28 12 |
15 12 13 31 13 |
15 12 13 31 13 |
| 14 12 14 29 10 |
14 12 13 31 10 |
14 12 13 31 10 |
| 14 14 14 30 13 |
17 14 13 29 10 |
14 12 13 29 10 |
| 17 13 13 31 13 |
14 12 12 29 13 |
14 12 14 28 10 |
| 17 12 13 30 13 |
14 13 13 29 13 |
16 12 13 29 13 |
| 14 12
13 31 13 |
|
|
|
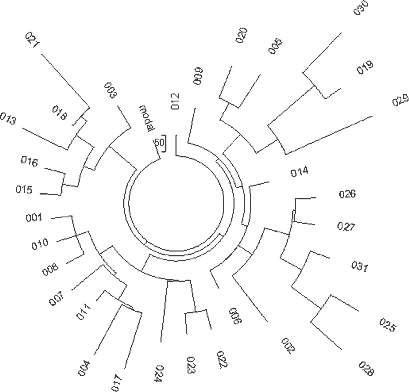
Figure 11. The 5-marker
haplotype tree for Rial haplotypes from China. The 31-haplotype tree was composed from
data provided by Dr. A.H.Bitttles and collected in ethnic communities
Hui (Uigurs), Bolan (Bao'an=Uigur),
Dongxi-ang (Bao'an=Uigur), and Sala
(Salar = Sary Uigur=Uigur)
(Bittles et al, 2007) (no haplotypes were provided in the referenced article). |
|---|
 |
|---|
234
These 31 5-marker haplotypes contain 99 mutations from the base haplotype
(in the FTDNA format)
13-X-14-X-X-X-X-12-X-13-X-30
which gives 0.639±0.085 mutations per
marker, or 21,000±3,000 years to a common ancestor. Such a large timespan to a common
ancestor results from a low mutation rate constant, which was calculated from the
Chandler’s data for individual markers, as 0.00677 mutation/haplotype/generation, or
0.00135 mutation/marker/generation (see Table 1 in Part I of the companion article).
Since haplotypes descended from such a ancient common ancestor have many mutations, which
makes their base (ancestral) haplotypes rather uncertain, the ASD permutational method
was employed for the Chinese set of haplotypes (Part I). The ASD permutational method
does not need a base haplotype and it does not require a correction for back mutations.
For the given series of 31 five-marker haplotypes the sum of squared differences between
each allele in each marker equals to 10,184. It should be divided by the square of a
number of haplotypes in the series (961), by a number of markers in the haplotype (5) and
by 2, since the squared differences between alleles in each marker were taken both ways.
It gives an average number of mutations per marker of 1.060. After division of this value
by the mutation rate for the five-marker haplotypes (0.00135 mut/marker/generation for 25
years per generation), we obtain 19,625±2,800 years to a common ancestor. It is within
the margin of error with that calculated by the linear method, as shown above.
It is likely that Haplogroup R1a1 had appeared in South Siberia around 20 thousand years
ago, and its bearers split. One migration group headed West, and had arrived to the
Balkans around 12 thousand years ago (see below). Another group had appeared in China
some 21 thousand years ago. Apparently, bearers of R1a1 haplotypes made their way from
China to South India between five and seven thousand years ago, and those haplotypes were
quite different compared to the Aryan ones, or the “Indo-European” haplotypes. This is
seen from the following data.
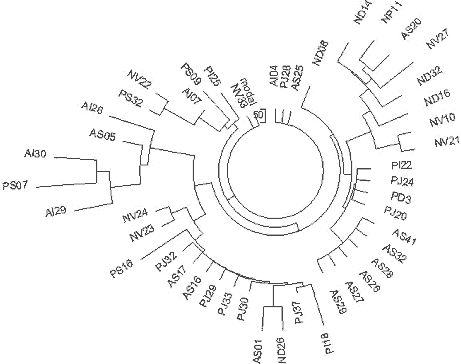
| Figure 12. The 6-marker haplotype tree for R1a1 haplotypes in Andra Pradesh (tribes
Naikpod, Andh, and Pardhan), South India. The 46-haplotype tree was composed from data
listed in (Thanseem et al, 2006). The designations of haplotypes are those used in the
article. |
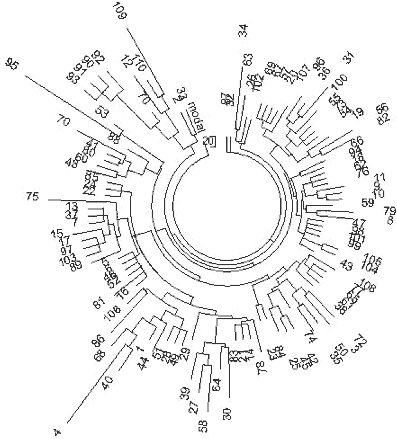
Figure 13. The 10-marker haplotype tree for R1a1 haplotypes in India (mixed
population, including tribes and castes). The 110-haplotype tree was composed from data
listed in (Sengupta et al, 2006). The article contains 114 Indian Rial haplotypes,
however, four of them were incomplete. |
 |
 |
235
Thanseem et al
(2006) found 46 R1a1 haplotypes of six markers each in three different tribal population
of Andra Pradesh, South India (tribes Naikpod, Andh, and Pardhan). The haplotypes are
shown in a haplotype tree in The 46 haplotypes contain 126 mutations, so there were
0.457±0.050 mutations per marker. This gives 7,125±950 years to a common ancestor. The
base (ancestral) haplotype of those populations in the FTDNA format is as follows:
13-25-17-9-X-X-X-X-X-14-X-32
It differs from the “Indo-European” Indian haplotype,
13-25-16-10-11-14-12-12-10-13-11-30
by four mutations on six markers, which corresponds
to 11,850 years between their common ancestors, and places their common ancestor at
approximately (11850+4050+7125)/2 = 11,500 ybp.
236
110 of 10-marker R1a1 haplotypes of various Indian populations, both tribal and Dravidian
and Indo-European castes, listed in (Sengupta et al, 2006) shown in a haplotype tree in
Figure 13, contain 344 mutations, that is 0.313±0.019 mutations per marker. It gives
5,275±600 years to a common ancestor. The base (ancestral) haplotype of those
populations in the FTDNA format is as follows:
13-25-15-10-X-X-X-12-10-13-11-30
It
differs from the “Indo-European” Indian haplotype
13-25-16-10-11-14-12-12-10-13-11-30
by
just 0.5 mutations on nine markers (DYS19 has, in fact, a mean value of 15.5 in the
Indian haplotypes), which makes them practically identical. However, in this haplotype
series two different populations, the “Indo-European” one and the “South-Indian” one were
mixed, therefore an “intermediate”, apparently phantom “common ancestor” was artificially
created with an intermediate TSCA, between 4,050±500 and 7,125±950 (see above).
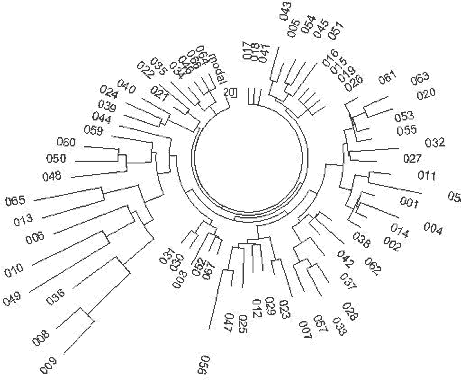
For a comparison, let us consider Pakistani R1a1 haplotypes listed in the article by
Sengupta (2003) and shown in Fig. 14. The 42 haplotypes contain 166 mutations, which gives
0.395±0.037 mutations per marker, and 7,025±890 years to a common ancestor. This value
fits within the margin of error to the “South-Indian” TSCA of 7,125±950 ybp. The base
haplotype was as follows:
13-25-17-11-X-X-X-12-10-13-11-30
It differs from the
“Indo-European” Indian haplotype by two mutations in the nine markers.
| Figure 14. Pakistani R1a1 |
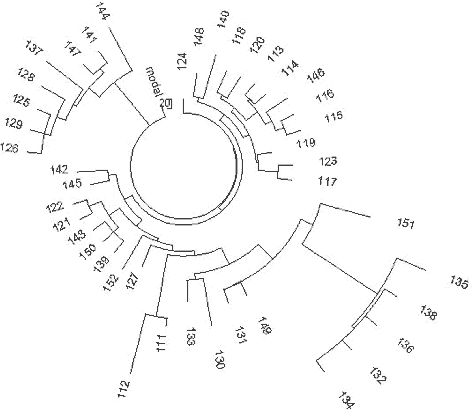
Figure 15. Balkan R1a1 |
 |
 |
Finally, we will consider Central Asian R1a1 haplotypes listed in the same paper by
Sengupta (2006). Ten haplotypes contain only 25 mutations, which gives 0.250±0.056
mutations per haplotype, and 4050±900 years to a common ancestor. It is the same value that we have
found for the “Indo-European” Indian haplotypes.
237
The above suggests that there are two different subsets of Indian R1a1 haplotypes. One
was brought by European bearers known as the Aryans, seemingly on their way through
Central Asia, in the 2nd millennium BCE, another, much more ancient, made its way
apparently through China, and arrived in India earlier than seven thousand years ago.
R1a1 Haplotypes of the Arabian Peninsula
Sixteen R1a1 10-marker haplotypes from Qatar and United Arab Emirates have been recently
published (Cadenas et al., 2008). They split into two branches, with base haplotypes
13-25-15-11-11-14-X-Y-10-13-11-30
13-25-16-11-11-14-X-Y-10-13-11-31
which are only slightly
different, and on DYS19 the mean values are almost the same, just rounded up or down. The
first haplotype is the base one for seven haplotypes with 13 mutations in them, on
average 0.186±0.052 mutations per marker, which gives 2,300±680 years to a common
ancestor. The second haplotype is the base one for nine haplotypes with 26 mutations, an
average 0.289±0.057 mutations per marker or 3,750±825 years to a common ancestor.
Since a common ancestor of R1a1 haplotypes in Armenia and Anatolia lived 4,500±1,040 and
3,700±550 ybp, respectively (Klyosov, 2008b), it does not conflict with 3,750±825 ybp in
the Arabian peninsula.
The Balkan Ancient Branch: the Oldest Trace of R1a Haplotype in Europe?
A series of 67 haplotypes of Haplogroup R1a1 from the Balkans was published (Barac et
al., 2003a, 2003b; Pericic et al., 2005). They were presented in a 9-marker format only.
The respective haplotype tree is shown in Fig. 15.
238
One can see a remarkable branch on the left-hand side of the tree which stands out as
an “extended and fluffy” one. These are typically features of a very old branch compared
with others on the same tree. Also, a common feature of ancient haplotype trees is that
they are typically “heterogeneous” ones and consist of a number of branches.
The tree in Fig. 15 includes a rather small branch of twelve haplotypes on top of the tree,
which contains only 14 mutations. This results in 0.130±0.035 mutations per marker, or
1,850±530 years to a common ancestor. Its base haplotype
13-25-16-10-11-14-X-Y-Z-13-11-30
is practically the same as that in Russia and Germany.
|
What happened 1850 years ago in E.Europe and impacted Germany, if not the influx of the
Hunnic tribes and creation of the Hunnic state? Are we looking at the haplotype that
describes the bulk of the Hunnic-circle people? |
The wide 27-haplotype branch on the right contains 0.280±0.034 mutations per marker,
which is rather typical for R1a1 haplotypes in Europe. It gives typical in kind 4,350±680
years to a common ancestor of the branch. Its base haplotype
13-25-16-11-11-14-X-Y-Z-13-11-30
is again typical for Eastern European R1a1 base haplotypes, in
which the fourth marker (DYS391) often fluctuates between 10 and 11. Of 110
Russian-Ukrainian haplotypes diagramed in Fig. 7, 51 haplotypes have “10”, and 57 have “11” in
that locus (one has “9” and one has “12”). In 67 German haplotypes, discussed above, 43
haplotypes have “10”, 23 have “11” and one has “12.” Hence, the Balkan haplotypes from
this branch are more close to the Russian than to the German haplotypes.
|
Seems to be another confirmation that the haplotype is connected with the influx of the
Hunnic tribes and creation of the Hunnic state. |
The “extended and fluffy” 13-haplotype branch on the left contains the following
haplotypes:
13 24 16 12 14 15 13 11 31
12 24 16 10 12 15 13 13 29
12 24 15 11 12 15 13 13
29
14 24 16 11 11 15 15 11 32
13 23 14 10 13 17 13 11 31
13 24 14 11 11 11 13 13 29
13 25
15 9 11 14 13 11 31
13 25 15 11 11 15 12 11 29
12 22 15 10 15 17 14 11 30
14 25 15 10 11
15 13 11 29
13 25 15 10 12 14 13 11 29
13 26 15 10 11 15 13 11 29
13 23 15 10 13 14 12 11
28
The set does not contain a haplotype which can be defined as a base. This is because
common ancestor lived too long ago, and all haplotypes of his descendants living today
are extensively mutated. In order to determine when that common ancestor lived, we have
employed three different approaches, described in the preceding paper (Part 1), namely
the “linear” method with the correction for reverse mutations, the ASD method based on a
deduced base (ancestral) haplotype, and the permutational ASD method (no base haplotype
considered). The linear method gave the following deduced base haplotype, an alleged one
for a common ancestor of those 13 individuals from Serbia, Kosovo, Bosnia and Macedonia:
13-24-15-10-12-15-X-Y-Z-13-11-29
The bold notations identify deviations from typical ancestral
(base) East-European haplotypes. The third allele (DYS19) is identical to the Atlantic
and Scandinavian R1a1 base haplotypes. All 13 haplotypes contain 70 mutations from this
base haplotype, which gives 0.598±0.071 mutations on average per marker, and results in
11,425±1,780 years from a common ancestor.
The “quadratic method” (ASD) gives the following “base haplotype” (the unknown alleles
are eliminated here, and the last allele is presented as the DYS389-2 notation):
12.92 –
24.15 – 15.08 – 10.38 – 12.08 – 14.77 – 13.08 – 11.46 – 16.62
A sum of square deviations
from the above haplotype results in 103 mutations total, including reverse mutations
“hidden” in the linear method. Seventy “observed” mutations in the linear method amount
to only 68% of the “actual” mutations including reverse mutations. Since all 13
haplotypes contain 117 markers, the average number of mutations per marker is
0.880±0.081, which corresponds to 0.880/0.00189 = 466±62 generations or 11,650±1,550
years to a common ancestor. 0.00189 is the mutation rate (in mutations per marker per
generation) for the given 9-marker haplotypes (companion paper, Part I).
A calculation of 11,650±1,550 years to a common ancestor is practically the same as
11,425±1,780 years, obtained with linear method and corrected for reverse mutations.
The all-permutation “quadratic” method (Adamov & Klyosov, 2008) gives 2,680 as a sum of
all square differences in all permutations between alleles. When divided by N2 (N =
number of haplotypes, that is 13), by 9 (number of markers in haplotype), and by 2 (since
deviation were both “up” and “down”), we obtain an average number of mutations per marker
equal to 0.881. It is near exactly equal to 0.880 obtained by the quadratic method above.
Naturally, it gives again 0.881/0.00189 = 466±62 generations or 11,650±1,550 years to a
common ancestor of the R1a1 group in the Balkans.
239
These results
suggest that the first bearers of the R1a1 haplogroup in Europe lived in the Balkans
(Serbia, Kosovo, Bosnia, Macedonia) between 10 and 13 thousand ybp. It was shown below
that Haplogroup R1a1 has appeared in Asia, apparently in China or rather South Siberia,
around 20,000 ybp. It appears that some of its bearers migrated to the Balkans in the
following 7-10 thousand years. It was found (Klyosov, 2008a) that Haplogroup R1b appeared
about 16,000 ybp, apparently in Asia, and it also migrated to Europe in the following
11-13 thousand years. It is of a certain interest that a common ancestor of the ethnic
Russians of R1b is dated 6775±830 ybp (Klyosov, to be published), which is about two to
three thousand years earlier than most of the European R1b common ancestors. It is
plausible that the Russian R1b have descended from the Kurgan archaeological culture.
|
It seems to indicate that the populations with old (20,000 ybp, R1a1) and new (16,000
ybp, R1b), and newer (11,500 ybp, newer “base” R1a1) never split, the migrating groups
carried a mixture of R1a1 and R1b, with their derivatives, around the Eurasia,
disseminating them on the way. The R1b modification that appeared in E.Europe 6,800 ybp
is associated with the Kurgan archaeological culture. |
The data shown above suggests that only about 6,000-5,000 ybp bearers of R1a1,
presumably in the Balkans, began to mobilize and migrate to the west toward the
Atlantics, to the north toward the Baltic Sea and Scandinavia, to the east to the Eastern
European Plains and steppes, to the south to Asia Minor, the Middle East, and far south to the
Arabian Sea. All of those local R1a1 haplotypes point to their common ancestors who lived
around 4,800 to 4,500 ybp. On their way through the Eastern European Plains and steppes the R1a1
tribe presumably formed the Andronovo archaeological culture, apparently domesticated the
horse, advanced to Central Asia and formed the “Aryan population” which dated to about
4,500 ybp. They then moved to the Ural mountains about 4,000 ybp and migrated to India as
the Aryans circa 3,600-3,500 ybp. Presently, 16% of the male Indian population, or
approximately 100 million people, bear the R1a1 haplogroup’s SNP mutation (SRY10831.2),
with their common ancestor of 4,050±500 ybp, of times back to the Andronovo
archaeological culture and the Aryans in the Eastern European Plains and steppes . The current
“Indo-European” Indian R1a1 haplotypes are practically indistinguishable from Russian,
Ukrainian, and Central Asian R1a1 haplotypes, as well as from many West and Central
European R1a1 haplotypes. These populations speak languages of the Indo-European language
family.
|
The split of haplotypes does not produce a new language, but a combination of disparate
groups and disparate languages does produce new languages. It appears that a mixture of
Old Europe language and R1a1 language produced the new *Proto-Indo-European language in
the Balkans ca 10,000 ybp. |
The next section focuses on some trails of R1a1 in India, an “Aryan trail.”
The Chenchu R1a1 Haplotypes
Kivisild et
al. (2003) reported that eleven out of 41 individuals tested in the Chenchu, an
australoid tribal group from southern India, are in Haplogroup R1a1 (or 27% of the
total). It is tempting to associate this with the Aryan influx into India, which occurred
some 3,600-3,500 ybp. However, questionable calculations of time spans to a common
ancestor of R1a1 in India, and particularly in the Chenchu (Kivisild et al., 2003;
Sengupta et al., 2006; Thanseem et al, 2006; Sahoo et al, 2006; Sharma et al, 2009;
Fornarino et al, 2009) using methods of population genetics rather than those of DNA
genealogy have precluded an objective and balanced discussion of the events and their
consequences.
The eleven R1a1 haplotypes of the Chenchus (Kivisild et al., 2003) do not provide good
statistics; however, they can allow a reasonable estimate of a time span to a common
ancestor for these 11 individuals. Logically, if these haplotypes are more or less
identical, with just a few mutations in them, a common ancestor would likely have lived
within a thousand or two thousands of ybp.
Conversely, if these haplotypes are all mutated, and there is no base (ancestral)
haplotype among them, a common ancestor lived thousands ybp. Even two base (identical)
haplotypes among 11 would tentatively give
ln(11/2)/0.0088 = 194 generations,
which,
corrected to back mutations, would result in 240 generations, or 6,000 years to a common
ancestor, with a large margin of error. If eleven of the six-marker haplotypes are all
mutated, it would mean that a common ancestor lived apparently earlier than 6 thousand
ybp. Hence, even with such a small set of haplotypes one can obtain useful and meaningful
information.
The eleven Chenchu haplotypes have seven identical (base) six-marker haplotypes (in the
format of DYS 19-388-290-391-392-393, commonly employed in earlier scientific
publications):
16-12-24-11-11-13
They are practically the same as those common East
European ancestral haplotypes considered above, if presented in the same six-marker
format:
16-12-25-11(10)-11-13
Actually, the author of this study, himself an R1a1 (Slav
R1a1) has the “Chenchu” base six-marker haplotype.
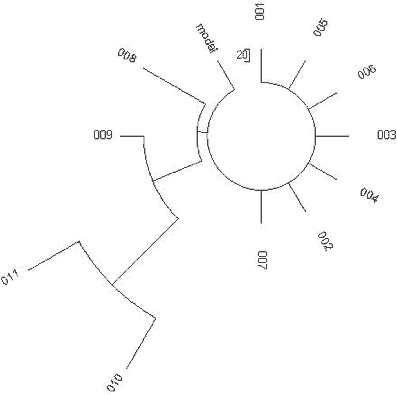
These identical haplotypes are represented by a “comb” in Fig 16. If all seven identical
haplotypes are derived from the same common ancestor as the other four mutated
haplotypes, the common ancestor would have lived on average of only 51 generations bp, or
less than 1300 years ago [ln(11/7)/0.0088 = 51], with a certain margin of error (see
estimates below). In fact, the Chenchu R1a1 haplotypes represent two lineages, one
3,200±1,900 years old and the other only 350±350 years old, starting from around the 17th
century CE. The tree in Fig 16 shows these two lineages.
240
| Figure 16. Chenchu R1a1 |
 |
A quantitative description of these two lineages is as follows. Despite the
11-haplotype series containing seven identical haplotypes, which in the case of one common
ancestor for the series, would have indicated 51 generations (with a proper margin of
error) from a common ancestor, the same 11 haplotypes contain 9 mutations from the above
base haplotype. The linear method gives 9/11/0.0088 = 93 generations to a common ancestor
(both values without a correction for back mutations). Because there is a significant
mismatch between the 51 and 93 generations, one can conclude that the 11 haplotypes
descended from more than one common ancestor. Clearly, the Chenchu R1a1 haplotype set
points to a minimum of two common ancestors, which is confirmed by the haplotype tree (Fig. 16). A recent branch includes eight haplotypes, seven being base haplotypes, and one with
only one mutation. The older branch, contains three haplotypes containing three mutations
from their base haplotype:
15-12-25-10-11-13
The recent branch results in ln(8/7)/0.0088
= 15 generations (by the logarithmic method), and 1/8/0.0088 = 14 generations (the linear
method) from the residual seven base haplotypes and a number of mutations (just one),
respectively. It shows a good fit between the two estimates. This confirms that a single
common ancestor for eight individuals of the eleven lived only about 350±350 ybp, around
the 17th century. The old branch of haplotypes points at a common ancestor who lived
3/3/0.0088 = 114±67 generations BP, or 3,200±1,900 ybp with a correction for back
mutations.
241
Considering that the Aryan (R1a1) migration to northern India took place about
3,600-3,500 ybp, it is quite plausible to attribute the appearance of R1a1 in the Chenchu by
3,200±1,900 ybp to the Aryans.
The origin of the influx of Chenchu R1a1 haplotypes around the 17th century is likely
found in this passage excerpted from (Kivisild et al., 2003): “Chenchus were first
described as shy hunter-gatherers by the Mohammedan army in 1694.”
Native American haplotypes of Haplogroup Q1a3a
(end of citation)
















