Ancient DNA provides new insights into the history of south Siberian Kurgan people
Christine Keyser • Caroline Bouakaze • Eric Crubézy • Valery G. Nikolaev • Daniel Montagnon • Tatiana Reis • Bertrand Ludes
Hum Genet (2009) Volume 126, p. 395–410, DOI 10.1007/s00439-009-0683-0 May 2009 ©
Critical Review
Links
Foreword
This posting attempts a critical review of the first comprehensive study of the Siberian Kurgan people, and a major contribution to the science of Turkology. The criticism is addressed to the historical aspects of the article, which technical results are admirable and significant, but were substantially complemented by A.Klyosov's analysis of R1a1 development, see The 3 R's in R1 Haplogroup. To dispel the claim of the authors who refer to Lamberg-Karlovsky as an unbiased arbiter on the Kurgan problem, his opinion (2005) is the opposite of what is asserted in this 2009 publication: "Given the increasingly large number of divisions and subdivisions of the generic Andronovo culture(s), with evidence for "no one group having undue prestige over the others," there is neither reason nor evidence to believe that they all shared an Indo-Iranian language. From the common roots of the millennia-long Andronovo culture(s) [and before that the related Timber Grave culture(s)], processes of both convergence and divergence [archaeologically indicated by the eastward migrations of the Andronovo culture(s)] allow for the presence of not only the Indo-Iranian languages but for other language families as well, that is, Altaic and Uralic. Clearly, the convergence of cultures, that is, the assimilation of local populations by an in-coming peoples, is very poorly developed within the archaeological discipline." A 2002 Lamberg-Karlovsky's statement reads: "There is, however, no compelling archaeological evidence that they (Andronovo and Bactrian Margiana archaeological complexes ) had a common ancestor or that either is Indo-Iranian." The content of the present study is exceedingly valuable; its spin method is an echo of the times discarded, and does a great disservice to the authors. A prejudiced analysis is not an analysis, but willful machinations, and nowadays fake references to authorities do not create facts on the ground.
The posting excluded main portions of the work that do not cause
objections, for a complete article please refer to the above link. From some excluded
paragraphs, only a pertinent phrase is cited, to provide a contextual background for the general comments. It
appears that the author's loose use of the word "European" is predominantly in a geographical sense, but
in places the authors clearly apply it to the Caucasoid phenotype that was linked with Cro-Magnon descent, a
Caucasoid phenotype marked by its "robust" features. The use of the term "European"
instead of the "Caucasoid" tends to cloud the subject, and was denoted in
this posting by replacing the inappropriate term with a
strikeout of the authors' misleading phrasing. The posting also denoted
some misleading phraseology peculiar to the Russian nationalistic propaganda, like
calling Ukraine and N.Pontic a southern Russia, and other oddities not picked up by
the publisher's editors. Among other peculiarities is a tradition of typological dating,
facilitated by the absence of instrumental dating, which clouds the study by
assigning specimens to a wide range from half a millennia to almost a millennia (e.g. 800 BC-100 AD).
Another shortcoming is an omission of the known anthropological background, connected
with cultural and genetical developments. The prior research noted changes in the
physique of the people, correlated with cultural changes, and the possible stages and
sources of the admixtures, which the genetical study is seeking to elucidate. The
Bronze Age kurgan cemeteries are known archeologically and anthropologically from Danube
in the west to the Kazakhstan and Siberia, and to Mongolia in the east, but genetical
studies so far are sorely missing, especially prominently so in the European Scythian area,
"southern Russia" in the lingo of the article, and
Kazakhstan. This spot-check study, being a very significant achievement, is only a peek
into a much larger and richer picture of the Eurasian steppe history. The affinity
between Germanic and Türkic people was noted
long ago, it is manifested in the ethnological traits, in culture, in mythology, in
linguistic affinity of the Germanic branch of the IE family and Türkic linguistic family,
and even in scripts, where Germanic and Türkic runes are two major entries among
the category of the runiform scripts, and this study adds a genetical facet to the common
traits, seemingly without even realizing it. The Karasuk people (S17-S20) were the first
who left archeological traces of milking cattle, which started widespread introduction of
the dairy products in the daily diets of the nomadic pastoralists, known well from
numerous classical sources that described that antic. Apparently, their genetics
overcome a lactose intolerance that is still widespread in the world, the lactose
genetics (C/T13910 at 2q21) was a useful tool for studies of the population movements in prehistoric
societies, i.e. precisely fitting the authors' objectives of tracing nomadic
pastoralists, and the absence of the lactose tolerance expressions from the design of the
investigation makes it much less then what it could be. We remain in the darkness whether
that trait was brought over from the Neolithic times by the Andronovo
Kurgan people, or developed within the Karasuk population.
For a professional repudiation by Anatole A. Klyosov of the authors' verbal and quasi-scientific equilibristic, go to http://www.lulu.com/items/volume_67/8049000/8049509/1/print/Vol2No5.pdf (Proceedings of the Russian Academy of DNA Genealogy, vol. 2, No. 5, 2009, pp. 871-878, in Russian).
The posting's comments and explanations added to the text of the article are shown in parentheses in (blue italics) or blue boxes.
Selected Citations
To help unravel some of the early Eurasian steppe migration movements,
we determined the Y-chromosomal and mitochondrial haplotypes and haplogroups of 26
ancient human specimens from the Krasnoyarsk area dated from between the middle of the
second millennium BC. to the fourth century AD. In order to go further in the search of
the geographic origin and physical traits of these south Siberian specimens, we also
typed phenotype-informative single nucleotide polymorphisms. Our autosomal, Y-chromosomal
and mitochondrial DNA analyses reveal that whereas few specimens seem to be related
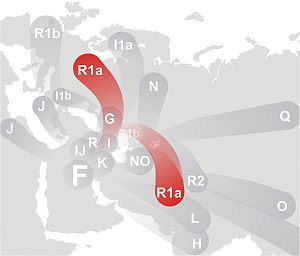
matrilineally or patrilineally, nearly all subjects belong to haplogroup R1a1-M17 which
is thought to mark the eastward migration of the Andronovo
Kurgan people early Indo-Europeans. Our results also confirm
that at the Bronze and Iron Ages, south Siberia was a region of overwhelmingly
predominant Caucasoid European
settlement, suggesting an eastward migration of Kurgan people across the
Eurasian Russo-Kazakh steppe.
Finally, our data indicate that at the Bronze and Iron Age timeframe, south Siberians
were blue (or green)-eyed, fair-skinned and light-haired people
(Ch. Di, Dinlin and Tele, Kirgizes) and that they might have played a role in the early
development of the Tarim Basin civilization. To the best of our knowledge, no equivalent
molecular analysis has been undertaken so far.
Kurgans (Türkic Russian
word for tumuli) are barrows (actually, cemeteries, with kurgans
being one of the physical attributes of a funeral ritual) characteristic of a culture arising on the
N.Pontic steppes of southern Russia
about 5000 BC and later spreading into eastern, central and northern Europe between 4400
and 2800 BC (and also into Middle and Central Asia, and Siberia). The Kurgan culture is divided into different sub-cultures on the basis of
the different kinds of graves under the barrows: Pit-Graves (Russ.
Yamna), Catacomb-graves (Russ. Katakomnaya) and
Timber-Graves (Russ. Srubna). The westwards diffusion
of this culture is sometimes equated with the appearance in eastern Europe of the Corded
Ware culture and the introduction of Indo-European-speaking peoples (Gimbutas 1970)
(Reference to Gambutas is inaccurate. The archeologist's Gambutas theory addressed
westward spread of the Kurgan
Culture, and advocated IE linguistic affiliation of the westward spread. Gambutas
herself, but not het theory, allowed participation of alternate linguistic phyla,
specifically Uralic and Türkic-Altaic. The archeologist Gimbutas linguistic analysis did
not extend eastwards of the N.Pontic, where Siberia happened to be. The archeologist Gambutas theory, which replaced previous
theories of Germanic invasions with the Baltic invasions has not been accepted in Russia as an official
doctrine, and in Soviet times has not been even officially acknowledged, which amounted
to a country-wide gag order, probably for racial-political
reasons. The Kurgan community united by common religious practices is dated by 5,000BC,
the beginning of the IE linguistic community is estimated at 4,000BC, and as all new
languages it had to be Creole language, born from the local languages. In that scenario,
the westward expansion of the IE in the 4,000BC started as a Creole, and the following
discrete waves were discrete modifications of that Creole. What happened in the
intervening millennia with the parental languages is anybody's guess, since the Gambutas
theory only follows the IE bud. History gives us all kinds of scenarios: Romans and
Germans vs. Romans and Judeans vs. Roman and Greeks; Germans and Balto-Slavs vs. Germans
and Gauls; Chinese and Huns vs. Chinese and Southern Huns vs. Chinese and Manchu; Slavs
and Bulgars vs. Slavs and Greeks vs Slavs and Germans, and so on. Anyone can pick up a
scenario to their liking, and build a convincing theory on it.)
In an attempt to reconstruct some of the population movements of ancient Kurgan people
within from the Eurasian steppes, the genetic background of 32 ancient human specimens from the
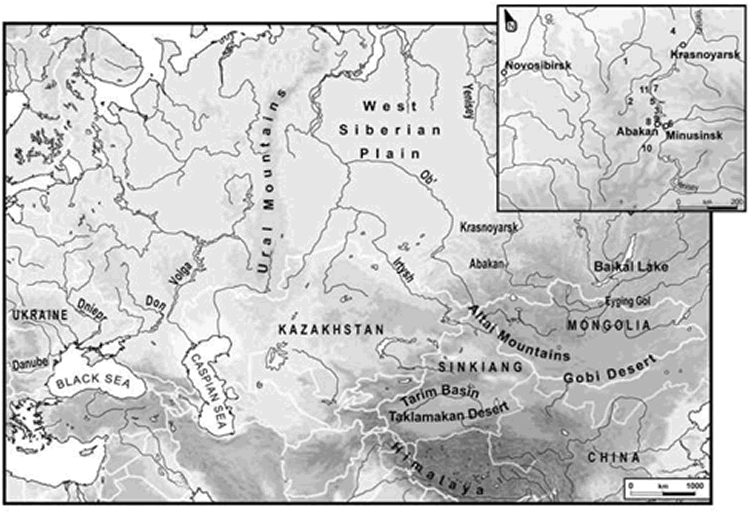
Krasnoyarsk area in southern central Siberia (along the Yenisey River; Fig. 1)
(commonly known as Minusinsk depression) was characterized at the nuclear and
mitochondrial DNA levels. Among these specimens, 10 were attributed to the Andronovo
culture, 4 to the Karasuk culture, 12 to the Tagar culture and 6 to the Tashtyk one
(Table 1). The Andronovo culture, related to the Timber-Grave group, appeared throughout
the south Russian steppe, Kazakhstan and western central Asia during the second
millennium BC. (Koryakova and Epimakhov 2007).
Numbers refer to the burial sites noted in Table 1
| Specimen | Code | Site/region/(map number) | Culture | Period | Dates | Sex |
|---|---|---|---|---|---|---|
| Bronze 1 | S07 | Tatarka cemetery, burial 64, Charypov district (1) | Andronovo | Middle Bronze Age | 1800-1400 BC | M |
| Bronze 2 | S08 | Tatarka cemetery, burial 55 Charypov district (1) | Andronovo | Middle Bronze Age | 1800-1400 BC | F |
| Bronze 3 | S09 | Solenoozernaia IV, kurgan I, burial 3 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | M |
| Bronze 4 | S10 | Solenoozernaia IV, kurgan I, burial 4 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | M |
| Bronze 5 | Sll | Solenoozernaia I, burial 4 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | F |
| Bronze 6 | S12 | Solenoozernaia I, burial 15 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | ? |
| Bronze 7 | S13 | Solenoozernaia IV, kurgan I, burial 4 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | ? |
| Bronze 8 | S14 | Solenoozernaia I, burial 4, Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | F |
| Bronze 9 | S15 | Solenoozernaia I, burial 29 Krasnoyarsk region (2) | Andronovo | Middle Bronze Age | 1800-1400 BC | ? |
| Bronze 10 | S16 | Oust-Abakansty, chief kurgan, Khakassia republic (3) | Andronovo | Middle Bronze Age | 1800-1400 BC | M |
| Karasuk 1 | S17 | Katcha, Drokino II, burial 1Emelyanov district (4) | Karasuk | Late Bronze Age | 1400-800 BC | M |
| Karasuk 2 | S18 | Oust-Abakansty, kurgan IV, burial 1Khakassia republic (3) | Karasuk | Late Bronze Age | 1400-800 BC | F |
| Karasuk 3 | S19 | Bogratsky, burial I, Khakassia republic (5) | Karasuk | Late Bronze Age | 1400-800 BC | F |
| Karasuk 4 | S20 | Minussinsk, Podgorny, burial 1 (6), | Karasuk | Late Bronze Age | 1400-800 BC | M |
| Tagar 1 | S21 | Novosselov district, Anach village, kurgan I, burial 3 (7) | Tagar | Iron Age | 800 BC-100 AD | ? |
| Tagar2 | S22 | Novosselov district, Anach village, kurgan II, burial 4 (7) | Tagar | Iron Age | 800 BC-100 AD | F |
| Tagar 3 | S23 | Tchernogorsk, burial 1, Khakassia republic (8) | Tagar | Iron Age | 800 BC-100 AD | ? |
| Tagar 4 | S24 | Tchernogorsk, burial 6, Khakassia republic (7) | Tagar | Iron Age | 800 BC-100 AD | M |
| Tagar 5 | S25 | Oust-Abakansty, Khakassia republic (9), | Tagar | Iron Age | 800 BC-100 AD | M |
| Tagar 6 | S26 | Bey district, burial 3, Khakassia republic (10) | Tagar | Iron Age | 800 BC-100 AD | M |
| Tagar 7 | S27 | Bograt district, kurgan 133, burial 3, Khakassia republic (11) | Tagar | Iron Age | 800 BC-100 AD | F |
| Tagar 8 | S28 | Bograt district, kurgan 133, burial 3, Khakassia republic (11) | Tagar | Iron Age | 800 BC-100 AD | M |
| Tagar 9 | S29 | Bograt district, kurgan 133, burial 3, Khakassia republic (11) | Tagar | Iron Age | 800 BC-100 AD | M |
| Tagar 10 | S30 | Bograt district, kurgan 133, burial 2, Khakassia republic (11) | Tagar | Iron Age | 800 BC-100 AD | ? |
| Tagar 11 | S31 | Bograt district, kurgan 133, burial 2, Khakassia republic (11) | Tagar | Iron Age | 800 BC-100 AD | ? |
| Tagar 12 | S32 | Bograt district, Abakano-Pérévoz II, burial 1, Khakassia republic (5) | Tagar | Iron Age | 800 BC-100 AD | M |
| Tashtyk 1 | S33 | Bograt district, Abakano-Pérévoz I, burial 5, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
| Tashtyk 2 | S34 | Bograt district, Abakano-Pérévoz I, burial 4, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
| Tashtyk 3 | S35 | Bograt district, Abakano-Pérévoz I, burial 2, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
| Tashtyk 4 | S36 | Bograt district, Abakano-Pérévoz I, burial 4, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
| Tashtyk 5 | S37 | Bograt district, Abakano-Pérévoz I, burial 4, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
| Tashtyk 6 | S38 | Bograt district, Abakano-Pérévoz I, burial 4, Khakassia republic (5) | Tashtyk | Iron Age | 100-400 AD | F |
Map number, noted in bracket, indicates the location of the grave on Fig. 1
The bearers of this Middle Bronze Age culture were strongly associated with the Indo-Iranians (this is utter nonsense from the Propaganda Ministry, in the framework of the Scytho-Iranian theory advocated by V.I.Abaev) and credited with the invention of the spoke-wheeled chariot (Lamberg-Karlovsky 2002) (Lamberg-Karlovsky not only does not advocate Scytho-Iranian theory, but raises objections to it. Reference to Lamberg-Karlovsky in respect to Indo-Iranians is inaccurate. Our French co-authors should have been a little more overt in dampening the IE enthusiasm of their Russian colleagues).
The Karasuk culture is a Late Bronze Age culture that succeeded the Andronovo culture in southern Siberia (late second millennium BC.). Karasuk people were farmers who practiced metallurgy on a large scale. They produced a realistic animal art, which probably contributed to the development of the later Scytho-Siberian animal art style (Karasuk culture is distinguished by Kipchak-type kurgan burials with stone tiling and with a rectangular fence of stone slabs staked vertically into the ground. Anthropologically, Karasuk people are Andronovo-type South Siberian "robust" Caucasoids, but culturally they were interacting toward the east. Karasuk people originated tamgas, monetary knifes, and milking of cattle. Karasuk people stratified into Scythian culture west from the Minusinsk territory, and a Hunnish culture to the east. The Karasuk culture's division was possibly reflected in a division of Pra-Türkic linguistic phylum into Ogur and Oguz branches. Cultural orientation to the east may have been connected with orientation of exogamous conjugal relations to the east, reflecting an eastern orientation of the political unions. The Karasuk culture coincides with propagation of a number of cultural innovations to China: horsebreeding, bronze, monetary knifes, mounted warfare. The Karasuk culture was the first positively identified nomadic culture that established allied relations with China in 636 BC. These relations did not come to an empty place, the date is the dating in the records; the first cultural dominance of the nomadic Kurgan Culture in China starts with the Yin stage of the Shang period, and dated a few centuries before the 1st millennium BC, which happen to coincide with the beginning of the European Scythian nomadic Kurgan Culture that emanated from the same place at the same time).
The Karasuk culture was replaced by the early Iron Age Tagar culture (first millenium
BC.) which flourished in Khakassia (southern part of the Krasnoyarsk Krai) producing an
art of animal motifs related to that of the Scythians of
N.Pontic southern European Russia.
On the Yenisey River, the Tagar culture was replaced by the Tashtyk culture, dating from the first to fourth century AD (Tashtyk culture is distinctly Hunnic culture).
To investigate the history and origin of these ancient Krasnoyarsk specimens, two uniparentally inherited marker systems were analyzed. Indeed, apart from giving information about paternal and maternal lineages, both the non-recombining portion of the Y-chromosome (NRY) and the mitochondrial DNA (mtDNA) have proven to be good indicators of migration events in human population history (Underhill and Kivisild 2007). Autosomal short tandem repeats (STRs) were also typed to confirm conventional sexing and to assess possible parentage relationships and/or exogenous contamination. Finally, since the specimens under study are thought to have been "Caucasoid" (Kozintsev et al. 1999; Lebedynsky 2003; Moiseyev 2006), phenotype-informative single nucleotide polymorphisms (SNPs) were also tested.
To widen the geographic scale of our study, we determined the Y-chromosomal haplogroup of several Hun (orig.:Xiongnu) specimens dated from the third century BC to the second century AD. Hun (orig.:Xiongnu) were Türkic nomadic tribes inhabiting the steppes north of China and controlling an empire stretching beyond the borders of modern-day Mongolia.
We also performed Y-SNP typing of one Scytho-Siberian specimen from the Sebÿstei site in the Altai Republic (Central Asia) dated from the middle of the fifth century BC. All these specimens were previously typed for autosomal and mtDNA polymorphisms (Keyser-Tracqui et al. 2003; Ricaut et al. 2004) (Ricaut et al result was 1 putative ancestral paleo-Asiatic population (read South Siberian) and 1 Chinese population).
Materials and methods
Ancient human samples
DNA was extracted from 32 ancient human skeletons excavated from different kurgan sites of the Krasnoyarsk region in southern Central Siberia during the years 1964-2000. In this area, the average temperature is 20°C below zero in winter. Even if the graves under the kurgans were not frozen at the excavation, in summer the temperature at the graves level is never over a few degrees Celsius above or below zero. After being listed and arranged in cardboard boxes, skeletal remains were sent to the Krasnoyarsk State Medical Academy of Russia, at Krasnoyarsk University, where they have been stored in a dry and cold environment. In 2004, the cardboard boxes were opened for sampling by two members of our team and transferred to Strasbourg, France, under appropriate storage conditions. On arrival in the laboratory, samples were frozen until DNA extraction to ensure their good preservation. The location of the kurgans is indicated on Fig. 1; their associated culture, the time period, the estimated age and the morphological sex of the specimens studied are presented in Table 1. Some skeletons came from the same kurgan (e.g., S27-S31); one of them was excavated from a chief’s kurgan (S16). Skeletal remains used for the DNA analyses were all long bone fragments. Moreover, the Y-SNPs typing was carried out on ten of the male Hun (orig.:Xiongnu) specimens of the Egyin Gol valley necropolis (Keyser-Tracqui et al. 2003) as well as on a Scytho-Siberian skeleton from the Sebÿstei site (SEB 96K2) found in a kurgan of the Altai Republic and dated from the middle of the fifth century BC. (Ricaut et al. 2004).
Autosomal STR analysis
The parentage relationships between individuals were tested by pairwise comparison of the profiles.
Y-chromosomal STR and SNP analysis
The STR haplotypes obtained were individually compared to the Y chromosome Haplotype Reference Database (YHRD) (http://www.yhrd.org) ( ~ 57,000 9-loci haplotypes as of December 2008 database search) as well as to a private world Y-STR database maintained by us and containing data retrieved from the literature ( ~ 38,000 9-loci haplotypes).
A set of 13 Y-chromosomal SNPs [M3, M9, M17, M45, M89, M173, M175, M216, M217, M242, 92R7, RPS4Y711 (M130), and Tat (M46)] characterizing Asian and Amerindian populations were also tested (references for SNP selection are given in Bouakaze et al. 2007).
The distribution of the different defined haplotypes among modern and ancient populations was investigated by exact sequence searches performed against ~ 40,400 -mtDNA haplotypes collected from the literature and maintained in a personal database.
Human pigmentation gene SNP analysis
Ten autosomal SNPs were selected because of either their association with normal human pigmentation variation, namely, eye, hair and skin color or their previously reported allele frequency differences between populations of the world.
Results
Autosomal STR analysis
Of the 32 individual remains analyzed by multiplex amplification, 6 DNA samples [S12, S17, S20, S30, S31 and S38 (Table 1)] appeared severely degraded. The remaining extracted samples gave 26 more or less complete allelic profiles. Consensus data are reported in Table 2.
Morphological and molecular typing results for sex determination were in accordance with each other except for two specimens (S09 and S34); nevertheless, since the DNA profiles obtained for these two individuals were almost complete, it is highly probable that molecular results were the correct ones. Furthermore, the amelogenin locus allowed us to deduce the sex of four specimens for which morphological indicators of sex were absent (S13, S15, S21 and S23).
Comparison of the profiles in pairs revealed no first degree relatives, even for specimens unearthed from the same kurgan or even burial. It should be noted, nevertheless, that 35% of the DNA profiles were incomplete which might have hampered the detection of close familial kinship (Only a blanket study like for Egyin Gol can produce systematic data on familial kinship).
Y-chromosomal STR and SNP analysis
To identify male lineages, an analysis of polymorphic markers located on the male-specific part of the Y-chromosome was performed. Seventeen Y-specific STRs were typed and used to construct haplotypes. All of the 10 male Siberian specimens were successfully typed at 14 loci (at least) and 6 different haplotypes were differentiated (Table 3). Three of them were shared between at least two specimens suggesting that some individuals could have belonged to the same paternal lineage (S10 and S16; S24, S34 and maybe S25; possibly S28 and S29) although buried in different kurgan and/or with no evidence of a close kinship link (cf., autosomal STRs).
| Specimen | Haplogroup | DYS 19 |
DYS 385 |
DYS 389I |
DYS 389II |
DYS 390 |
DYS 391 |
DYS 392 |
DYS 393 |
DYS 437 |
DYS 438 |
DYS 439 |
DYS 448 |
DYS 456 |
DYS 458 |
DYS 635 |
YGATA |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| S07 | C(xC3) | 15 | 12/13 | 14 | 30 | 22 | 9 | 12 | 14 | 14 | 10 | 11 | 19 | 15 | 16 | 22 | 11 |
| S10/S16 | R1al | 16 | 11/14 | 14 | 32 | 25 | 11 | 11 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | 12 |
| S24/S34 | R1al | 17 | 11/14 | 13 | 31 | 24 | 11 | 11 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | 13 |
| S25 | R1al | - | 11/14 | 13 | 31 | 24 | 11 | 11 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | - |
| S26 | R1al | 16 | 11/14 | 13 | 31 | 24 | 11 | 11 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | 13 |
| S28 | R1al | 16 | 11/14 | 14 | 31 | 25 | 11 | 11 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | 12 |
| S29 | R1al | - | 11/14 | 14 | 31 | 25 | 11 | 11 | 13 | 14 | 11 | - | - | 16 | 15 | 23 | 12 |
| S32 | R1al | 17 | 11/14 | 13 | 31 | 24 | 11 | 12 | 13 | 14 | 11 | 10 | 20 | 16 | 15 | 23 | 13 |
Pairwise comparisons of the haplotypes showed that except for S07, all the ancient male specimens bore closely related allelic profiles, differing at most at six loci (on the 17 tested) and always by one-step mutation only.
Haplogroup (hg) assignment, based on the Y-chromosomal SNP typing, revealed that except for S07, which was found to belong to hg C(xC3), all ancient specimens were affiliated to hg R1a1. This finding is in agreement with the relative similarity of the haplotypes mentioned above. While hg C has a distribution generally limited to populations of northern Eurasia, eastern Eurasia, Oceania, and the Americas, R1a1 is widely spread across Eurasia. It is found among western Eurasian, southern Asian, central Asian and Siberian populations. This haplogroup is thought to trace the migration patterns of the early Indo-Europeans, perhaps stemming from the Kurgan culture (Zerjal et al. 1999; Semino et al. 2000) (Actually, it was thought initially to be early Indo-Europeans, with some questionable non-Indo-European oddities, but with a wider range of studies, it became clear that the pattern has Indo-Europeans only as a special case).
The additional analysis performed on Hun (orig.:Xiongnu) specimens revealed that whereas none of the specimens from the Egyin Gol valley bore this haplogroup, the Scytho-Siberian skeleton from the Sebÿstei site exhibited R1a1 haplogroup.
A search in the YHRD database as well as in our own databank revealed that none of the Y-STR haplotypes obtained from the south Siberian samples perfectly matched (at 17 loci) those included in the databases. Nevertheless, when not all loci were scored, matches were found for all samples except two (S07 and S32) for which even the search based on the 9-loci minimal haplotype was fruitless (Table 4). The S10/S16 haplotype matched the most frequent R1a1 haplotype (12 loci) seen in the south Siberian population of Derenko et al. (2006). This haplotype is notably found at high frequency in Altaians. It carries an allelic stucture 16-14-32-25-11-11-13 (DYS19-DYS389I-DYS389II-DYS390-DYS391-DYS392-DYS393) which is considered as a founder haplotype relative to southern Altaians (Southern Altaians are essentially Tuvans, per L.Potapov) (Kharkov et al. 2007). The S10/ S16 haplotype is also found in eastern Europe (Hungary, Slovenia, Poland) as well as in Asia (Central Anatolia) (All these had a Türkic admixture).
Table 4 Results of the search against Y-STR databases(Without a relative frequency in each location, the informative value of the listing has a most generalized nature, especially so because of its dependence on the state of the statistics in each location, which are profoundly different; it should be a major factor in assessing the results)
| Sample | Minimal haplotype | Minimal haplotype +2 to 7 additional loci |
|---|---|---|
| S07 | No match | No match |
| S10/S16 | 1 Hungarian 2; 2 Slovenians, 2 Poles (YHRD) | 2 Tuvinians, 14 South Altaians 6; 1 Turk 4; 1 Hungarian 7; 1 Pole 22; 1 Cretan 16 |
| S24/S34 (S25) | 5 Poles 17, 19; 1 German 19; 2 Poles, 1 Nepalese, 1 Turk, 1 German, 1 Armenian (YHRD) | 1 German 20; 1 Pole 24; 1 Indian 8 |
| S26 | 1 German 10; 2 Poles 17, 19; 1 Serbian 23; 1 Pole, 1 Kazakh, 1 Nepalese, 2 Germans, 1 Turk (YHRD) | 1 Turk 4; 1 Chinese 15; 1 Indo-Pakistani 1; 8 South Siberians 6; 1 Indian 5; 5 Poles 24; 1 German 20 |
| S28(S29) | 2 Lithuanians 13, 19; 4 Latvians 13, 19; 6 Poles 17, 19, 1 German 19; 1 Byelorussian 18; 1 Russian 14; 2 Swedish 9, 11; 3 Serbians 23; 1 Estonian 12, 1 Lithuanian 12; 6 Slovenians, 13 Czechs, 4 Hungarians, 7 Germans, 3 Slovaks, 3 Ukrainians, 2 Croats, 1 Siberian Tuvan, 3 Poles, 3 Norwegian (YHRD) | 1 Austrian 2; 1 Lithunian 18; 1 Uigur 26; 20 South Siberians 6; 3 Poles 22; 2 Malays 25; 2 Indians 5; 2 Russians 21 |
| S32 | No match | No match |
|
The S24/S34 haplotype is mainly found in Poland and Germany (In the Late Antique time, territories of both Poland and eastern Germany had significant Türkic presence, which may explain their falling into a single class. Poland had a considerable influx of Türkic people in the Middle Age, but Germany did not.) In Asia it is found in Anatolia, Armenia, Nepal and India (Except for Nepal, three others had significant Türkic presence).
Haplotype of specimen S26 has a wide distribution since it appears in Europe as well as in western Asia, in Central Asia, in southern Asia and in southern Siberia. The allelic structure 16-24-11-11-13 (DYS19, DYS390, DYS391, DYS392, DYS393) found in this haplotype was described as the most frequent motif observed in a Ukrainian population by Kravchenko et al. (2002). According to these authors, this 5 Y-STR-loci haplotype might be an ancestral one (Ukraine was a permanent home of a number of Türkic populations).
Haplotype S28 is the most frequently found in present-day populations. It is essentially carried by eastern and northern Europe individuals, as well as south Siberians (Footprint coincides with European Sarmatia and European Huns).
The S32 haplotype was not found in the databases even though it differs from the S24/S34 haplotype by only one-step mutation at locus DYS392.
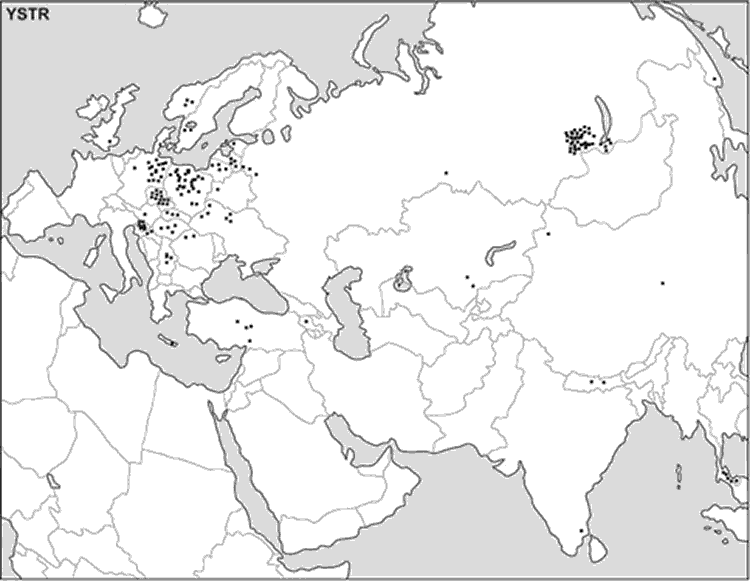
The S07 haplotype also did not appear in the YHRD database even when one mismatch was allowed in the minimal haplotype search. The current distribution pattern of all the Y-STR haplotypes found in our ancient sample is reported in the map on Fig. 2.
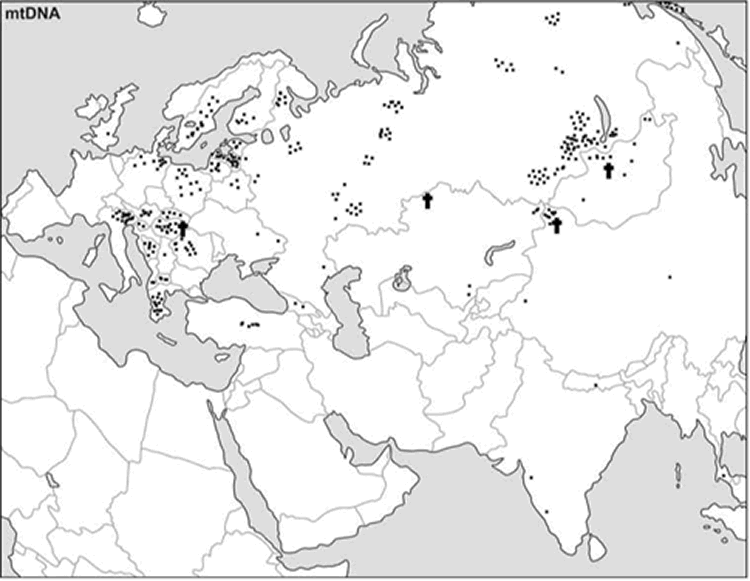
Fig. 2 Current distribution pattern of the Y-STR haplotypes found in the
ancient Siberians under study.
Each square represents a present-day individual sharing
the same Y-haplotype of an ancient specimen
(The vast white areas on the map do not necessarily indicate
that no traces of the Kurgan People exist in those areas, they rather indicate that no
data compatible with the findings of this study exist in the databases. Nowhere in the
description is noted the present spotty character of the databases used for comparisons,
the potential consequences of the future additions to the databases, and the prospective
areas for future studies. The difference between mtDNA map and Y-DNA map may simply
indicate that for the territories of the former Soviet Union, data bases contain more
spot studies of the mtDNA than the Y-DNA. Ditto for Mongolia, and Türkic, Mongol, Manchu,
and Tungus minorities in China. Without these qualifications, the analysis may present
seriously distorted conclusions)
Mitochondrial DNA analysis
The mtDNA haplotype of the 26 ancient individuals for whom genotypes were obtained, was determined by sequencing of the HVI region. To avoid ambiguous conclusions and to corroborate haplogroup assignment, all individuals were additionally typed for some specific mtDNA coding SNPs. Results are indicated in Table 5. A good agreement was found between coding and control region data except for three samples (S27, S29, S35) sharing a CRS HVI haplotype whose SNapShot assay coding SNPs allowed classification as hg U. Moreover, the SNapShot assay allowed us to obtain additional information regarding the phylogenetic refinement of two samples, S13 and S32, found to belong to sub-hgs H6 and H5a, respectively. Overall, 23 different haplotypes were distinguished and assigned to 16 different haplogroups. Twenty samples were found to belong to west Eurasian haplogroups (U2, U4, U5a1, T1, T3, T4, H5a, H6, HV, K, and I), whereas the 6 remaining samples were attributed to east Eurasian haplogroups (Z, G2a, C, F1b and N9a).
Table 5 HVI haplotype and haplogroup attribution for each Krasnoyarsk specimen successfully analyzed and current distribution of the haplotypes| Sample | HVI haplotype | MtHg | SNPHg | Occurrence of the haplotype in the world |
|---|---|---|---|---|
| S07/S14 | 356C | U4 | - | 2 Turks 7, 38; 6 Russians 1, 19, 33; 1 Ukrainian 33; 5 Mansi 10; 18 Volga-Ural region individuals 2; 2 Germans 40, 48; 17 Altai-Sayan region individuals 12, 18, 46; 5 Bosnians 34; 1 Slovenian 34; 2 Uygur 54; 1 Uzbek 54; 3 Mongols 54, 26, 14; 2 Chechens 42; 15 Latvians 39, 31; 1 Albanian 3, 4 Macedonians 3, 57, 3 Romanians 3, 4 Hungarians 4, 22; 1 Buryat 14; 1 Finn 20; 3 Poles 19; 1 Lithuanian 31, 3 Seto 31, 3 Karelians 31, 3 Swedish 31; 4 Slovaks 35; 6 Greeks 23; 3 Italians 49; 1 ancient Hungarian 50 |
| S08 | 129A 185T 223T 224C 260T 298C | Z(Z1) | Z | 2 Mongols 28, 29; 1 Kazakh 8; 8 Koryaks 45, 1
Itel'men 45; 1 Russe 33; 1 Ket 11, 2 Nganasans 11, 52; 3 Yukaghirs 52; 11 Volga-Ural region individuals 2; 1 Buryat 14, 1 Altaian 14, 2 Teleuts 14; 1 Volot 19, 2 Karelians 31, 1 Swedish 31 |
| S09 | 126C 163G 186T 189C 294T | Tl | Tl | 2 Turks 8,
12 Italians 17, 49; 1 Chinese Han 55; 1 Indian 0;
1 Uzbek 54; 5 Latvians 31, 39; 1 Mongol 26; 8 Hungarians 4, 22, 50; 3 Austrians 5; 3 Altaians 14
1 Chukchi 14; 4 Finns 20; 3 Germans 48; 2 Greeks 28; 3 Byelorussians 1^ Estonians 31, 4 Lithuanians 31, 1 Seto 31, 5 Karelians 31, 8 Swedish 31; 1 ancient Kazakhstan 30 and 1 ancient Eastern Turkistan (orig.: Xinjiang) 17 specimens |
| S10 | 051G 092C 129C 183C 189C 362C | U2e | U2 | 1 Estonian 31; 1 Uygur 54 |
| Sll | 126C 294T 324C | T4 | T | 2 Roma 24; 1 Byelorussian 1,! Hungarian 22; 1 Greek 23 |
| S13 | 187T362C | H | H6 | Corsican 16 |
| S15 | 093C224C311C319A | K2b | UK | 1 Hungarian 4; 1 Austrian |
| S16 | 192T 256T 270T | U5al | U5al | 1 Russian 1; 2 Kets 11; 1 Byryat 14; 1 Khakassian 14; 1 Mongol 26; 1 Albanian 3;
2 Macedonians 3; 3 Romanians 3, 4 Greeks 23; 3 Hungarians 4, 22; 2 Austrians 5, 2
Italians 49; 1 Finn 20, 1 Lithuanian 31, 2 Swedish 31 |
| S18 | 114A256T270T294T | U5al | U5al | 1 Northwestern European 44 |
| S19 | 356C 362C | U4* | U4 | 1 Indian 43; 1 Hungarian 4 Volga-Ural region individuals ; 2 Altai—Sayan region individuals 3;
1 Estonian 31, 1 Lithuanian 31; 1 Bosnian 34, 2 Slovenians 34; 1 Austrian 5; 1 Greek 23; 2 Italians 49 |
| S21/S22 | 126C 189C292T294T | T3 | T | 2 Sardinians 15 |
| S23 | 126C 189C 292T 294T 296T | T3 | T | Not found |
| S24 | 129A223T304C391A | 1(14) | - | 1 Indian 27; 2 Icelanders 21; 1 Buryat 14; 5 Swedish 31; 5 Hungarians 22; 1 ancient Scandinavian 36 |
| S25 | 093C 223T 227G 278T 362C | G2a | G2 | 1 Uygur 54; 3 Koreans 14, 29, 41; 2 Latvians 31, 39 |
| S26 | 148T 223T 234T 288C 298C 327T | C | C | 1 Tuvinian 14 |
| S27 | CRS | H | u | Too frequent |
| S28 | 172C 179T 183C 189C 232A 249C 304C 311C | Fib | Fl | 2 Mongols 14, 54 |
| S29 | CRS | H | U | Too frequent |
| S32 | 304T319A | H(H5) | H5a | 1 Austrian 5 |
| S33 | 093C 129A 223T 298C 327T | C | C | 2 Kazakhs 8, 54; 6 Altaians 13; 7 Buryats 13, 46;
15 Tuvinians 13, 46; l Uygur 54; 2 Chineses 53, 41; 10 Evenks 14, 461 1 Ulchi 46; 6 Yakuts 47, 14; 2 Mongols 14, 411 1 Kalmyk 141 2 Oroqen 41; 1 Hun (orig.: Xiongnu) 25 |
| S34 | 172C311C | HV | HV | 1 Indian 43; 1 Uzbek 54, 1 Mongol 54; 1 Evenk 41; 1 Kalmyk 14; 1 Macedonian 57; 2 Italians 49, 51 |
| S35 | 093C 209C | H | U | 1 Estonian 31 |
| S36 | 223T 257A 261T | N9a | N9a | 5 Chinese 41, 53, 551 3 Koreans 32, 411 56; 2 Vietnameses 23 |
| S37 | 126C 163G 186T 189C 269G 294T 362C | Tl | Tl | Not found |
To assess the present distribution of the mtDNA types found in the ancient sample, a search of their occurrence among modern populations of Eurasia was carried out in the literature and personally compiled database. Exact matches were observed for almost all sequences.
Among those mostly prevalent was the hg U4 motif 356C (S07/ S14) which was found in northern, eastern and southeastern European populations, as well as in Volga-Ural, Altai-Sayan and peri-Baikal area populations. Note that this sequence occupies a central position on U4 phylogeny built by Malyarchuk (2004). It was also observed in an ancient Hungarian specimen from the tenth to eleventh century (Tömöry et al. 2007). The sub-hg U4 variant 356C-362C (S19) is present in northern, eastern and Mediterranean Europe, in the Volga-Ural and Altai-Sayan regions as well as in southern India (All heavily Türkic-populated areas).
Another sequence within hg U showing a wide geographic distribution is the S16-U5a1 haplotype which matches in southern Siberia, in Central Asia, as well as in northern, western, eastern and southeastern Europe.
Conversely, the S18-U5a1 haplotype is most likely rare since found only once in a northwestern European, likewise the S10-U2 haplotype which has been described in one eastern European and one central Asian individual only.
Specimens S27, S29 and S35 bear a CRS HVI sequence belonging to hg U. Such results have already been described in Russian and Byelorussian populations where hg U CRS sequences were found in a relatively high proportion ( ~ 35%; Belyaeva et al. 2003).
Within hg T, the most prevalent sequence type was that harbored by specimen S09. This sequence was described as the root sequence of hg T1 (Richards et al. 2000; Pike 2006). Its highest frequency is in western Eurasia (mainly the Baltic region) with occasional occurrences in eastern Eurasia. This founder haplotype was also observed in two ancient specimens, one from Kazakhstan (1400-1300 BC, Bronze Age; Lalueza-Fox et al. 2004), the other from Eastern Turkistan (orig.:Xinjiang) site in northwestern China (Gao et al. 2008).
Surprisingly, the S37-haplotype, differing from the previous one by two additional mutations, was not observed in our database.
The T3-haplotypes harbored by specimens S21, S22 and S23 are either rare or absent whereas the S11-T4 haplotype, although being described as a main founder cluster within haplogroup T (Richards et al. 2000), was not commonly found.
The H haplotypes observed in our ancient sample are uncommon in present-day populations since both S13-H6 and S32-H5a sequences were found in only one European individual. HV lineage are represented by one sequence (S34) found in India, in Central Asia and in southeastern Europe.
Specimen S24 was found to belong to haplogroup I, subclade I4. Exact matches to this I4 type were mainly found in northern and eastern Europe individuals. The fact that one ancient Scandinavian specimen (0-400 AD) bore this sequence gives direct evidence of the antique presence of such sequence in the north of Europe (Melchior et al. 2008) (Recall the Scandinavian runic inscriptions unreadable in Germanic, but readable in Türkic).
The K2 haplotype harbored by specimen S15 was observed only twice, in European samples (1 Hungarian and 1 Austrian) (Hungary/Austria harbored generations of ethnically various Türkic tribes).
The eastern Eurasian lineages are represented by sequences belonging to hgs N9a, Z, G2a, F1b and C. The Z haplotype observed in the S08 ancient specimen belong to subhg Z1. It is observed in northeastern Asians, in south Siberian populations as well as in Central Asia. It is also present among several populations of the Volga-Ural and Baltic Sea regions.
The S25-G2a sequence has been observed in few but dispersed individuals (Koreans, Latvians and Uygur).
The specimen S28 belongs to hg F1b with a motif reported in two Mongols only (Mongolia had Türkic admixture at least from the 2nd c. BC).
Haplogroup C is represented by two sequences: one had a HVI motif observed in only one south Siberian individual (S26). The other had a HVI motif mainly found in Siberia and Central Asia (S33).
Finally, the S36 specimen carries a N9a-haplotype identical to those described previously in East Asian individuals (Chinese, Koreans, Vietnamese).
Figure 3 represents the current and past distribution of the overall mtDNA haplotypes found in our ancient Krasnoyarsk sample (except the CRS sequences). This distribution is similar to that depicted in Fig. 2 for the Y-chromosome, despite the sparser pattern of the mtDNA counterpart (Which points to mass migrations by whole families, as opposed to an army).
Human pigmentation gene SNP analysis
In order to deepen the search of the geographic origin of the Siberian specimens under study, we typed SNPs located in human pigmentation genes. Ten SNP markers located in genes that have been described as accounting for variation in human hair, eye and skin color but also in ethnogeographic ancestry were thus selected and a minisequencing-based assay was developed on modern samples (Bouakaze et al. 2009). This assay was subsequently applied on the ancient Siberian samples so that complementary information provided by phenotype-associated SNPs could add to previous anthropological and genetic findings. The phenotype and ancestry of the ancient Siberian specimens under study are indicated in Table 6 (genotype details for each investigated marker is given in Bouakaze et al. 2009).
| Sample | Eye color according to rs12913832 |
Phenotype according to OCA2 diplotype |
Ancestry according to rs1545397, rs 16891982, rs2031526 |
|---|---|---|---|
| S07 | Brown | Blue or brown eye, dark brown hair, fair or medium skin | Asian |
| S08 | Brown | - | Asian |
| S09 | Blue | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S10 | - | Blue or brown eye, brown hair | European |
| S11Bl | Blue | - | European |
| S13 | - | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S14 | Brown | Blue or brown eye, brown hair, fair or medium skin | European |
| S15 | Blue | - | European |
| S16 | Blue | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S18 | Blue | - | European |
| S19 | Blue | - | European |
| S21 | Blue | Blue or brown eye, brown hair, fair or medium skin | European |
| S22 | Brown | - | European |
| S23 | Blue | - | European |
| S24 | Blue | - | European |
| S25 | Blue | - | European |
| S26 | Brown | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S28 | Brown | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S29 | Blue | - | European |
| S32 | Brown | - | Asian/European Mix |
| S33 | Blue | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S34 | Blue | - | European |
| S35 | Brown | - | European |
| S36 | Blue | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
| S37 | Blue | Blue or brown eye, blond or light brown hair, fair or medium skin | European |
- indicates that the marker could not be amplified
(All these Europeans, Asians, and Asian/European Mixes have the same light pigmentation of eyes, hair, and skin. Show me a single pure Indian or Iranian blue-eyed blond, and I will take you to a puppet show absolutely for free. Mind you, the Indo-Iranian does not have to come from from India, Iran, or even Ossetia, any place of migration will be gladly accepted, as long as the genes are purely Indo-Iranian)
Surprisingly, the typing of a SNP associated to eye color (rs12913832) shows that at least 60% (15/25) of the Siberian specimens had blue (or green) eyes (S27 cannot be tested because bone sample and DNA extract were used up). Moreover, the pigmentation SNP analysis showed that all except three specimens exhibited a European ancestry (Europe must have received a pigmentation patent from the Supreme Command), even when they bore an Asian mtDNA haplotype as is the case for samples S25, S26, S28, S33 and S36, demonstrating the importance of studying both maternal and paternal lineages. These results also show that two individuals carrying the same mtDNA haplotype can be classified in opposite ethnogeographic groups as is the case for samples S07 and S14: note that these two specimens belong to different paternal lineages since S07 is the single specimen carrying haplogroup C(xC3) but not haplogroup R1a1. Specimen S32 appears as having a mixed ancestry; curiously, this specimen exhibits Y-chromosome and mtDNA haplotypes virtually unknown in present-day populations. Most of the specimens seem to have been light-skinned people with blond or light brown hair (Exactly in accordance with what the Chinese annals are saying. What a real surprise for a scholar!).
|
Fig. 3 Current distribution pattern of the mtDN A haplotypes found in the ancient Siberians under study (except the CRS sequence). Each square represents a present-day individual sharing the same mtDNA haplotype of an ancient specimen. Cross represents an ancient specimen different from those studied in this work |
 |
Discussion
With the present study, we aimed to unravel some of the early Eurasian steppe migration movements by analyzing paternal, maternal and autosomal genetic variation in Bronze and Iron Age anthropological remains recovered from the Krasnoyarsk area in southern Central Siberia. There is virtually no knowledge either about the origins and the history of these ancient south Siberian inhabitants or about the language(s) they may have spoken. Nevertheless, many scholars believe that these Kurgan people, and notably the bearers of the Andronovo culture, spoke a Proto-Indo-Iranian or a Proto-Iranian language (Lamberg-Karlovsky 2002) (Lamberg-Karlovsky asserts that the beliefs of "many scholars" are based on beliefs, not on evidence). Moreover, the south Siberian tribes under study (Andronovo, Karasuk, Tagar) have been described as exhibiting pronounced Europoid (i.e. Caucasoid) features (Kozintsev et al. 1999; Lebedynsky 2003; Moiseyev 2006). These data raised questions as to where these people came from, which routes have been followed and to which extent they have contributed to the spread of the Indo-European language (or any other potential language, for that matter; quite the opposite, no IE speakers descended from the Siberian Kurgan cultures, as should have been noted in a nonpartisan discussion, the IE fell on them both in the prehistorical and historical times). In an attempt to answer these questions, we analyzed, in parallel, markers of distinct genetic systems.
Twenty-six out of the 32 bone samples collected yielded amplifiable DNA and 65% of the genetic profiles were complete (Table 1). This high success rate suggested that the nucleic acids were well preserved and allowed us to envisage other single-copy nuclear genes analyses (i.e., pigmentation genes). No close kinship link was detected between the subjects under study even for those found in the same kurgan (S27-S31). Nevertheless, the Y-chromosomal analyses performed on the ten male specimens showed that S28 and S29 shared the same haplotype. These Y-chromosomal analyses, based on the combined use of STRs and SNPs, also revealed that, with the exception of one individual, all samples examined fall into hg R1a1.
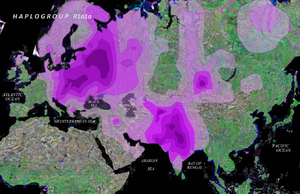
Haplogroup R1a1 is defined by marker M173 plus M17 (Y Chromosome Consortium 2002; Jobling and Tyler-Smith 2003; Karafet et al. 2008) and has a widespread distribution area on the Eurasian continent. It is spread among western Eurasian (mostly eastern European and Volga-Ural populations) (which are specifically known as being Türkic lands and the territory of the Türkic Kipchak Khanate for half a millennia; the present population is divided into Türkic substrate in the south, and Finnic substrate in the north), southern Asian (mainly India and Pakistan's populations) (which are specifically known as being Türkic-dominated lands for the duration of the Mughal period), central Asian (predominantly Türkic for two millennia) and Siberian populations (especially southern Siberians) (predominantly Türkic until the 20th c.), whereas it is rather rare in East Asian populations (together with associated traits of light hair/eye pigmentation and lactose tolerance). In western Eurasia, a clear northeast/south-west cline has been described (Rosser et al. 2000; Semino et al. 2000; Wells et al. 2001). Indeed, the R1a1 haplogroup frequency reaches a maximum in Poland, Hungary, and Ukraine and decreases in the direction of central and northern Europe (together with associated traits of light hair/eye pigmentation and lactose tolerance). The same occurs in the southern direction, towards Anatolia and the Caucasus. These clinal frequency distributions have been associated with ancient population movements in Europe: according to Semino et al. (2000), the geographical distribution of the R1a1 haplogroup probably reflects the re-population of Europe after the last glacial maximum ( ~ 20-12 kya) from a refugium in eastern Europe, likely in Ukraine (Passarino et al. 2001). This postglacial spread might have been magnified by the movement of the Kurgan people from the north of the Caspian Sea in a much more recent timescale (Rosser et al. 2000; Semino et al. 2000). This "Kurgan people" expansion would have resulted in the spread of the Indo-European language as postulated by Gimbutas (1970) (or non-IE languages, as was also admitted by the archeologist Gimbutas). Thereby, R1a1 was viewed by some authors as the marker of the Indo-European contribution (Zerjal et al. 1999; Kharkov et al. 2004). According to Pericic et al. (2005), the present distribution pattern of the R1a1 haplogroup was probably also influenced by much later migratory events like massive Slavic migration from fifth century AD (who at the time were ruled by Türkic Huns, Avars, Bulgars, intermingled with them, and spoke Turkified dialects of the Baltic languages. The omission of the very historical Huns, Avars, Bulgars, Khazars, Magyars, Badjanaks, Oguzes, Kipchaks, Mongolo-Tatar Kipchaks, that changed the face of the Eastern and Central Europe, and enormously affected the demography and genetical composition of these areas, is notoriously and glaringly missing from the official Russian historiography, and makes a mockery of a scientific discourse. It appears that propagation of intentional blindness has not abated in the beginning of the third millennia. The accepted norms of scientific and personal decency would require abandoning the blinds and demonstrate a a smattering of objectivity).
In our ancient sample, among the nine specimens carrying haplogroup R1a1, five different Y-chromosomal haplotypes were observed. Similarities were noted between these haplotypes, particularly the motif 11/14-11-11-13-14-11-10-20-16-15-23 (in bold in Table 3) which is common to all of them except S32. This motif is typically an eastern European one since currently found in the Russian federation only (YHRD database) (which would tend to point to the Akathyrs and Bulgars as the most demographically powerful populations of the early Eastern Europe). Matching haplotypes were found for all the R1a1-specimens except S32. Figure 2 shows that the current distribution pattern of the Y-STR haplotypes found in our ancient sample resembles that of R1a1. Indeed, they were observed at high frequencies in Slavic and Baltic populations (with peaks among Poland and Czech Republic) as well as in the indigenous populations of south Siberia. By contrast, they were only sporadically observed in central and east Asia and were absent in western Europe (which is consistent with the known historical events of the last 2 millennia).
Regarding the mtDNA analyses, our findings indicate that the ancient Krasnoyarsk mtDNA pool harbored both western and eastern Eurasian lineages. Nevertheless, most of the retrieved sequences (n = 20, 77%) belong to western Eurasian mtDNA haplogroups (HV, H, T, I, U and K). The eastern Eurasian lineages (23% of the sequences) were represented by haplogroups or subhapologroups C, Z, G2a, F1b and N9a. The western Eurasian contribution to the ancient mtDNA pool reached 90% for the Bronze Age and decreased to 67% for the Iron Age. Thus, despite a small sample size, our data suggests a temporal pattern which is in agreement with the view that west Eurasian populations predominated in the Krasnoyarsk region during the Bronze Age, whereas Asian component began to increase from the Iron Age on. This result is similar to that obtained in the ancient DNA study of Lalueza-Fox et al. (2004) who showed that all Kazakh sample specimens from before thirteenth to seventh centuries BC belonged to European lineages. After that time, there was an influx of East Asian sequences which are thought to have coexisted with the prior west Eurasian genetic substratum (which is consistent with the reconstructed archeological pre-history, and known historical events of the last 2 millennia).
As shown in Table 5, and particularly in Fig. 3, the current
distribution of the ancient mtDNA haplotypes can be broadly divided into three different
geographic poles. The first is represented roughly by eastern and northern Europe, the
second by the Volga-Ural region and the third by southern Siberia. It is interesting to
note that the distribution of the paternal and maternal lineages is close. Indeed, except
for the Volga-Ural region, both maps overlap. This would mean that the story of women
matches well that of men (and brings about a new valuable
insight that the Volga-Ural region was invaded by military expeditionary forces which
used local women for procreation, which is consistent with the Bulgarian historical
memories of the arrival of the Huns at the turn of the eras, and their takeover of the
control of the territory that persisted until their dynastic line was extinguished in the
6th c. AD, and replaced with a Dulo dynasty. . Notably, the tribal symbols of the Dulo clan of the Bulgars and the Qayn (Kayı) tribe of the Oguzes are the same
![]() ). In other words, the
(2 out of 3 major) migrations in which south Siberian specimens were
involved seemed to be "whole-population movements" rather than "war-like movements"
involving the men only.
). In other words, the
(2 out of 3 major) migrations in which south Siberian specimens were
involved seemed to be "whole-population movements" rather than "war-like movements"
involving the men only.
The fact that East Asian mtDNA sequences appeared at the Iron Age could signify that once settled, migrants of supposed European ancestry (i.e. anthropologically and genetically Caucasoid ancestry) began to establish relationships with groups coming from the east and to take Asian women as wives. Moreover, the relative high diversity of the mtDNA gene pool observed in the ancient specimens indicates that numerous populations carrying different mtDNA variants were involved in the formation of southern Siberian populations, even reflecting long-distant movements. It would not have presented any major difficulty for Bronze Age and Early Iron Age peoples to range from one end of Eurasia to the other within some centuries. Historical records and archaeology attest that nomadic groups moved across Eurasia from North of the Black sea, through Central and Inner Asia, to northeast Asia in a matter of centuries (Mair 2005). Some of them are described in Chinese historiography as horse-riding, Caucasian-looking, Indo-European-speaking people (allusion that Chinese described Hu, Juns, and Huns as Indo-European-speaking people is, naturally, totally false) and are sometimes referred as the "Kurgan Culture" (Zerjal et al. 2002) (for a simple reason that they buried their dead in kurgans). Paleogeographic studies provide material which suggests that climate change, particularly in the eastern regions of the steppes, was among the causes of these population movements (Van Geel et al. 2004) (in addition to the genocidal Chinese campaigns of divide and rule so vividly described in the Chinese annals).
If we consider that there is a correspondence between the overall distribution of haplotypes and haplogroups and past human movements, it seems that the European or Caucasoid component observed in the ancient Siberian sample may originate from East European populations. Moreover, it is likely that some mtDNA lineages were carried to southern Siberia from the Volga-Ural region. Incidentally, in the fifth century BC, Herodotus mentioned transit trade occurring in Central Asia along a route that stretched from the Urals in the west to the Altai and the Minusinsk Basin in the east (Hemphill and Mallory 2004). In Altai, the presence of the R1a1 haplogroup in the middle of the fifth century BC is confirmed by the sample SEB 96K2 of Ricaut et al. (2004) which was found to belong to this Y-haplogroup. The boundary of the eastern European influence seems to be fixed at the peri-Baikal area since no R1a1 haplogroup was found in the Hun (orig.:Xiongnu) specimens of the Northern border of Mongolia (For mobile nomadic people this absence of R1a1 haplogroup and any boundaries are nonsense, see for example K.Kim 2010).
According to the "Kurgan hypothesis" of Marija Gimbutas, nomadic peoples of the Volga steppe region, assumed to speak a Proto-Indo-European language, infiltrated Europe in three waves between 4400 and 2800 BC. Around 4400 BC, Kurgan people from the lower Dnieper and lower Volga regions began moving along the Black Sea littoral into the Danube Basin. They migrated in the Central Balkans and further into Central Europe. During the middle of the fourth millennium BC, the Kurgan culture in the North Pontic Region continued to develop. People travelled across western Ukraine north of the Carpathian Mountains to Poland and Central Germany. They also moved southwest into eastern Romania. Shortly after 3000 BC, the third Kurgan wave (Pit-Grave; orig.: Yamna people), originating once more from the Volga steppe, spread from Central Europe to Northwest Germany, the east Baltic area, southern Scandinavia, the upper Dnieper basin and Central Russia. These three waves of migrations might explain the distribution of mtDNA and Y-chromosome lineages observed in the present work (Figs. 2, 3) (that explanation would be a real Houdini-type achievement, since Gimbutas largely discredited theory of invasion waves brings Kurgans westward to Europe, not eastward to the Siberia of the present work. For the westward expansion, the consensus is that it was predominantly peaceful and symbiotic, which allowed deep penetration of the migrant's culture and innovations into the local diverse populations of the Central Europe and Balkans. Following the examples of the later expansions during the historical periods, of which we know hundreds, all of them, excluding those that joined kindered populations, lost their languages and now speak Scottish and English (Picts, Alans), Romanian (Scythians, Huns, Bulgars), Slavic (Badjanaks, Bulgars, Oguzes, many others), Hungarian (Scythians, Huns, Avars, Bulgars), Indian (Indo-Scythians, Kushans, Ephtalites, Mughals), Persian (Parthians, Mongols), Pashtu and Dari (Tochars, Ases, Sibirs, Ephtalites), Chinese (Huns, many others), Nakh (Tochars, Ases), Mongol (Huns), Armenian (Sakasena, Bulgars), Georgian (Kipchaks), Scandinavian (Sarmatians, Huns) and generally speaking any other language that borders the Eurasian steppes. Chances that the migrants imposed their language on the local populations are statistically non-existent. The study of Altaisms in the surrounding languages is in its incipience, but even the initial results are loud and clear, just recall that the Slavo-Turkic language of the "Tale of Igor's campaign" transitioned from the initial indignation and sensationalism to a thriving industry).
The Andronovo culture was preceded by the Afanasievo one, which is held to share the closest similarities with the (Pit-Grave; orig.: Yamna) culture found in the Pontic-Caspian region (Hemphill and Mallory 2004). An eastward migration of the (Pit-Grave; orig.: Yamna)-derived Afanasievo populations in the Eastern steppe thus provides a possible explanation for the appearance of a European component (i.e. Cro-Magnon descent, a "robust" Caucasoid phenotype of the eastern European Pit-Gravers) in the gene pool of ancient south Siberians (do we have an understanding what was the gene pool of ancient south Siberians before the arrival of the Afanasievo migrants? In science, a change is a delta of before and after, if the delta is non-existent, the change is zero, and a presumption of change is speculative).
Whereas archaeological records are inconclusive about the anthropological traits characteristic of ancient Siberians, our data deduced from the analysis of human pigmentation gene SNPs seems consistent with the fact that most of them had blue (green) eyes. Indeed, among the SNPs tested was rs12913832, a single DNA variation within a regulatory element of HERC2 gene which is associated to blue eye color in humans. This polymorphism, together with the diplotypes obtained from variations of the OCA2 locus (major contributor to the human eye color variation) showed that at least 60% of the ancient Siberian specimens under study had blue (or green) eyes. Such color phenotype is, according to Eiberg et al. (2008), caused by a founder mutation which most likely originated 6-10 kya from a region around the Black sea, near modern-day Ukraine or Turkey and then diffused into Northern Europe (i.e diffused into Northern Europe also, along with the non-Northern Europe Northern Eurasia. The speculation about its origin, one would think, is rested on the results of the studies, instead of presumptions, and has a solid explanation on the events that wiped out that trait from the same Ukraine or Turkey). Our data also suggest that south Siberian specimens might have had blond or light brown hair and fair skin and that they were of European ancestry, a result which appears as evident as those of uniparental markers ( this is a logical absurdity; Europe is predominantly populated by non-blonds, this rhetoric expands a particular trait, peculiar to the non-Indo-European Finno-Ugrians and European geographical zones under their influence, to the Europe as a whole, making it a fake innate ethno-geographical marker. The blond Spaniards, Italians, Greeks, Albanians, Iranians, Indians, Arabs, etc. may exist only in popular Hollywood movies, and even then they are limited to the main heroes. The Gauls, Balts, Dacians, Scandinavians were not brunettes, but heir connection with 6-10 kya Ukraine or Turkey is more then tenuous. This kind of logic serves for evidence in scientific speculations).
Interestingly, the haplotype of specimen S09 matches that of an ancient specimen from the Yuansha site (Takla-makan desert, Eastern Turkistan (orig.:Xinjiang Province), northwestern China) and dated back to 2,135 ± 50 years (Gao et al. 2008), suggesting genetic relationships between Andronovo populations and those of the Eastern Turkistan (orig.:Xinjiang). The Bronze Age inhabitants of the Eastern Turkistan (orig.:Xinjiang) were intrigued at their "Caucasoid" physical appearance and putative "European" origins (Mallory and Mair 2000) (in this case, the interviewed Bronze Age "inhabitants" of the Eastern Turkistan were Chinese military colonists, and the nomads of the Takla-makan in question were the Eastern Huns, comprised of the Hun and Uigur tribes, whose descendents remain at present distinct from Chinese by their "Caucasoid" physical appearance, but nothing of the putative "European" origin). Two hypotheses have been offered by archaeologists to account for the origins of these Bronze Age people (i.e of the historical Hun and Uigur tribes whose ancestral ranges included Takla-makan steppe surrounding Takla-makan oases, well documented in the Chinese annals for the period between the 2,185 BC and 2,085 BC of Gao's investigation. The 26,000 oasis agricultural settlers were a mix of the neighboring brunette Punjab agriculturists with their much more powerful and numerous nomadic Tele blond neighbors known at the time under a politonym Hun) believed to have spoken an Indo-European language called Tocharian (i.e. Kuchean and Hotan vernaculars mislabeled by first European explorers as Tocharian) and depicted as possessing red or blonde hair, long noses and blue or green eyes: the "steppe hypothesis" and the "Bactrian oasis hypothesis". Proponents of the latter assert that settlement of the Eastern Turkistan (orig.:Xinjiang) came from sedentary based population of the Oxus civilization found in Uzbekistan, Afghanistan and Turkmenistan (who, being a mix of the eastern and western branches of the Pit-Grave people are as distinct as, say, Dutch from Holland and Aruba), whereas proponents of the "steppe hypothesis" maintain that the Tarim region experienced a colonization attributed to Afanasievo and Andronovo populations who migrated to Eastern Turkistan (orig.:Xinjiang) from the Altai-Minusinsk regions north of the Tarim Basin (who were the eastern branch of the same Pit-Grave people, and would carry exactly the same biological, and practically identical cultural traits) (Hemphill and Mallory 2004). Our results corroborate the "steppe hypothesis" (but only for the case if the Siberian component of the "Oxus civilization" is ignored, and a total biological isolation between Siberian and Amu-darya branches is presumed, which totally contradicts the archeological determination of extensive cultural exchange between these two areas).
An essential aspect of the present work is the confidence in the validity of our data.
To conclude, in this work we demonstrated that some carriers of the Kurgan culture, believed to be Indo-European speakers (by some, but far from all scholars), were also carriers of the R1a1 haplogroup. These data lend further support to the idea that R1a1 might be a marker to the migration patterns of the early Indo-Europeans, an idea also supported by the recent article of Haak et al. (2008) in which individuals of the Corded Ware Culture, a culture commonly (i.e. frequently) associated with Indo-European, might bore R1a1 Y-chromosome (as we deduced from their Y-STR typing results) (The link between the Corded Ware Culture and the nomadic culture in the east is a long-established fact; that the local population spoke languages that belong to the Indo-European family may be conjured, as well as alternates, but assigning the Indo-European language to the eastern migrants without the Gimbutas' violent invasion continues to be a matter of undying controversy among the Indo-Europeanists, for reasons that the castle was built without evidentiary foundation).
The modern distribution of lineages is the outcome of many millennia of population movements and therefore the assumption of a Proto-Indo-European speaker's homeland in Kurgan region should be taken with great caution. Nevertheless, our study opens possibilities for new debates. We also showed for the first time (first only in the genetical code sense, because otherwise that fact was well known since the first archeological explorations) that Bronze and Iron Ages south Siberian populations displayed "European" (i.e. "Caucasoid"; the quotation marks around "European" should have been used not only in the punch line, but throughout the article to avoid perception of intentional manufacturing of scientific reality) physical appearance, thus corroborating physical anthropological records. Another conclusion that can tentatively be inferred from the data presented here is that the Andronovo culture might be the eastern spread of the Kurgan culture and might be related to Tocharian speakers in the Tarim Basin (in all fairness, the quotation marks also belong to the adjective "Tocharian").